Help Center
Comprehensive guide to using the PRIME platform for Polycomb regulatory research
Navigation
What is PRIME?
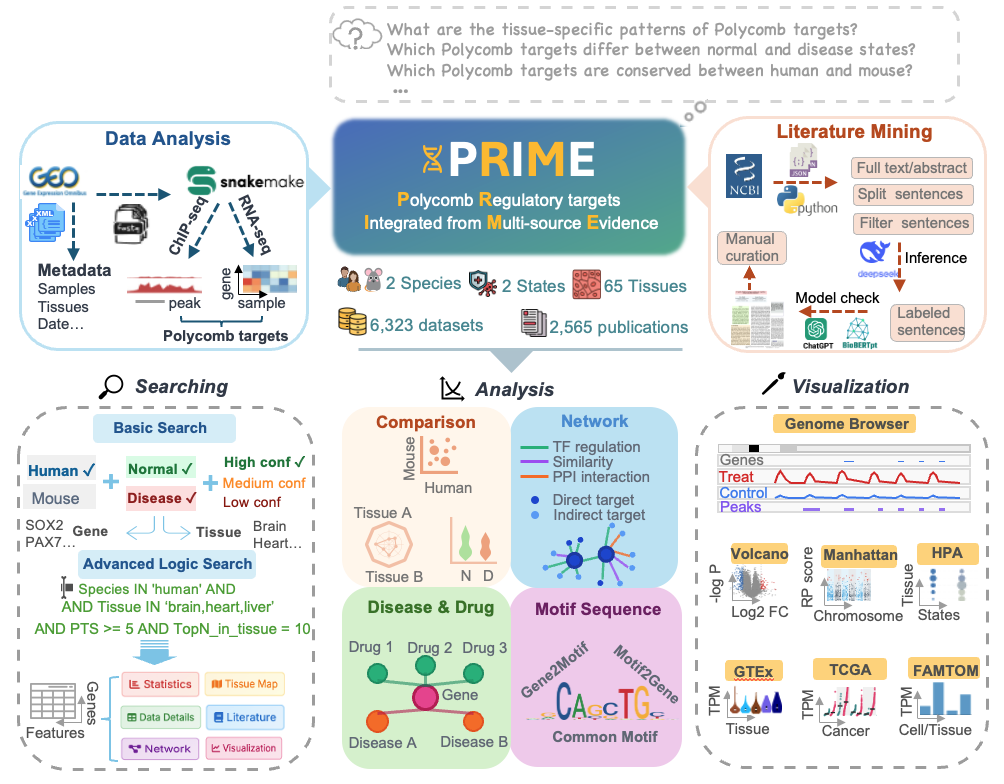
PRIME (Polycomb Regulatory targets Integrated from Multi-source Evidence) is a comprehensive scientific database and web platform for exploring Polycomb regulatory complex targets across multiple species and tissue types. The platform integrates multi-omics data including ChIP-seq, RNA-seq, and literature evidence, providing researchers with advanced search, analysis, and visualization capabilities for publication-quality Polycomb research.
- Comprehensive multi-omics integration: Incorporates 5,905 H3K27me3 ChIP-seq and 418 RNA-seq datasets, 5,381 literature-derived associations, and integrates external resources such as GTEx, TCGA, HPA, and FANTOM5, covering 65 tissue types under normal and disease conditions for both human and mouse (hg38/mm10).
- Data reliability: Employs a unified, standardized analysis pipeline with stringent quality control across all datasets. An original, multi-dimensional weighted scoring system integrates ChIP-seq regulatory strength, RNA-seq response, and cross-dataset consistency, enabling robust classification of associations into high, medium, or low confidence.
- Advanced visualization and analysis: Features a comprehensive suite of interactive tools and modules, including dynamic plots (volcano, MA, Manhattan), expression distribution visualizations (violin, dot, bar plots), multi-panel statistical dashboards, high-resolution figure export, an integrated genome browser, and interactive platforms for cross-dataset comparisons, regulatory network visualization, disease-drug associations, and motif analysis.
How to Search?
PRIME supports three search methods to find Polycomb targets:
1. Category Search
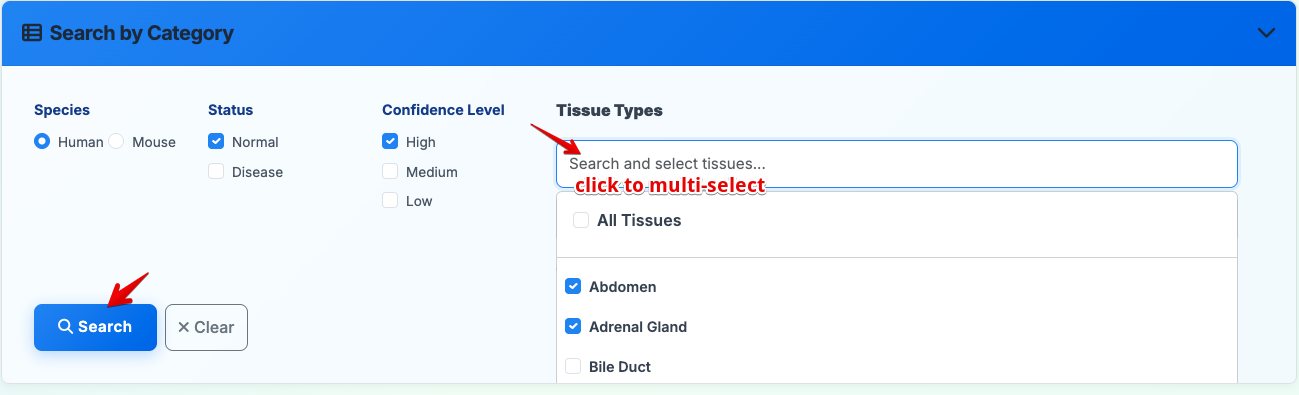
Search by tissue, status, species, and confidence level:
- Species: Select Human or Mouse
- Tissue Types: Choose one or multiple tissues (default: all tissues)
- Sample Status: Filter by normal or disease status (multi-select supported)
- Confidence Level: Set to High, Medium, or Low (multi-select supported)
- Execute: Click Search to display filtered results
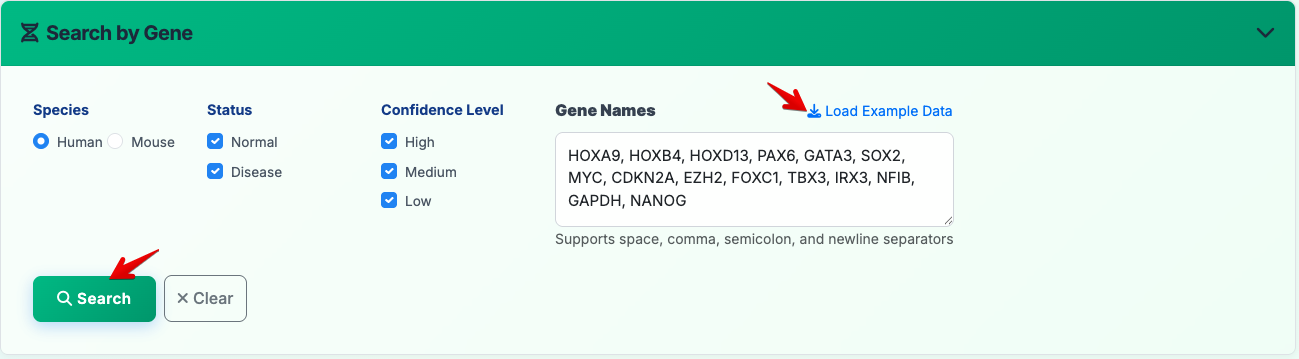
2. Gene Search
Search by gene names:
- Species: Select Human or Mouse (radio button selection)
- Status: Filter by Normal and/or Disease samples (checkbox selection)
- Confidence Level: Choose High, Medium, and/or Low confidence results (checkbox selection)
- Gene Names: Enter gene symbols in the text area (Click "Load Example Data" for quick testing with sample gene lists)
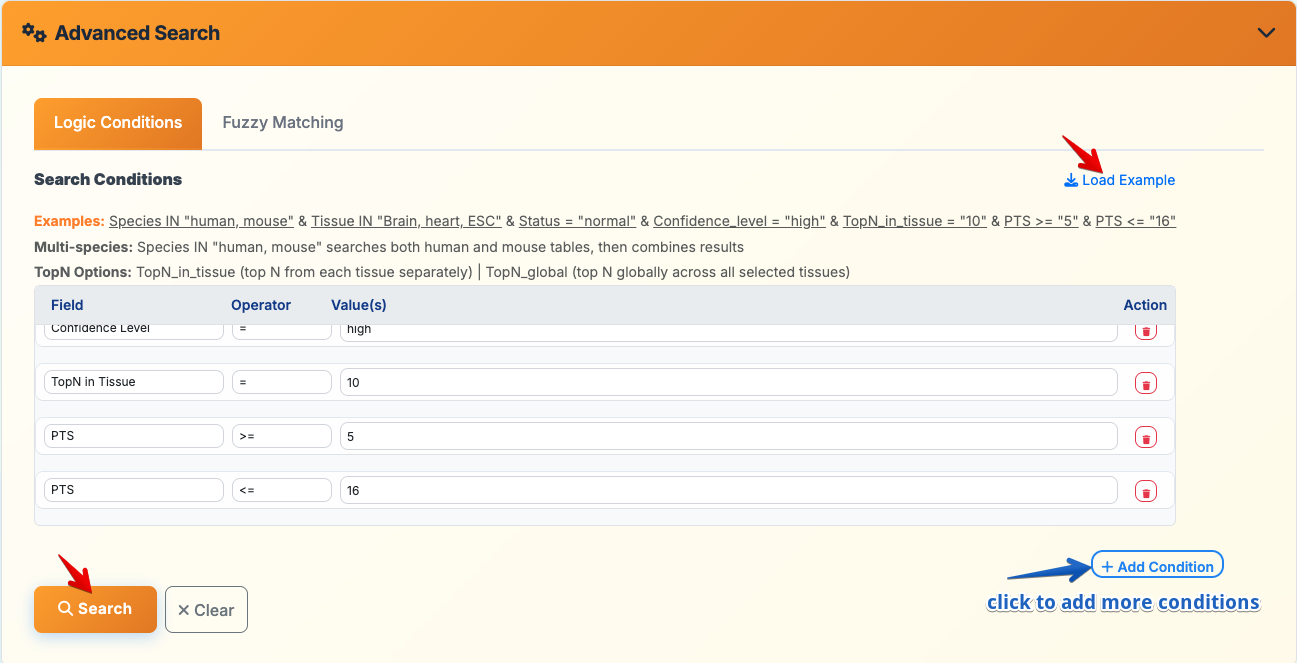
3. Advanced Search
Logic Conditions Mode:
- Flexible Query Builder: Add multiple search conditions using field-operator-value combinations
- Supported Fields: Species, Tissue, Status, Confidence Level, TopN settings, PTS values, etc.
- Multi-species Support: Search across human and mouse databases simultaneously
- TopN Options:
- TopN_in_tissue: Top N results from each tissue separately
- TopN_global: Top N results globally across all selected tissues
- Example Query:
Species IN "human, mouse" & Tissue IN "Brain, heart, ESC" & Status = "normal" & Confidence_level = "high"
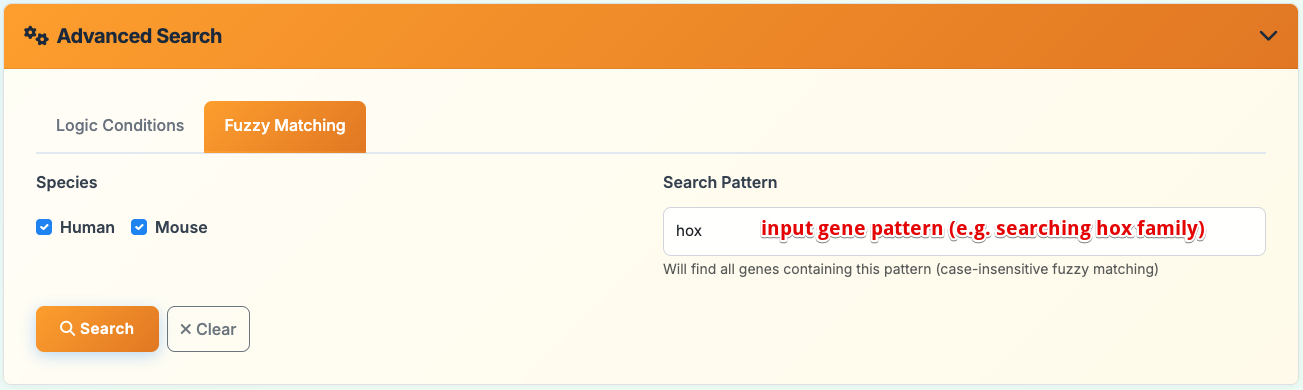
Fuzzy Matching Mode:
- Species Selection: Choose Human and/or Mouse (checkbox selection)
- Pattern Search: Enter gene name patterns for case-insensitive fuzzy matching
- Search Examples: hox, pax, sox (finds all genes containing these patterns)
- Flexible Matching: Identifies genes with partial name matches across selected species
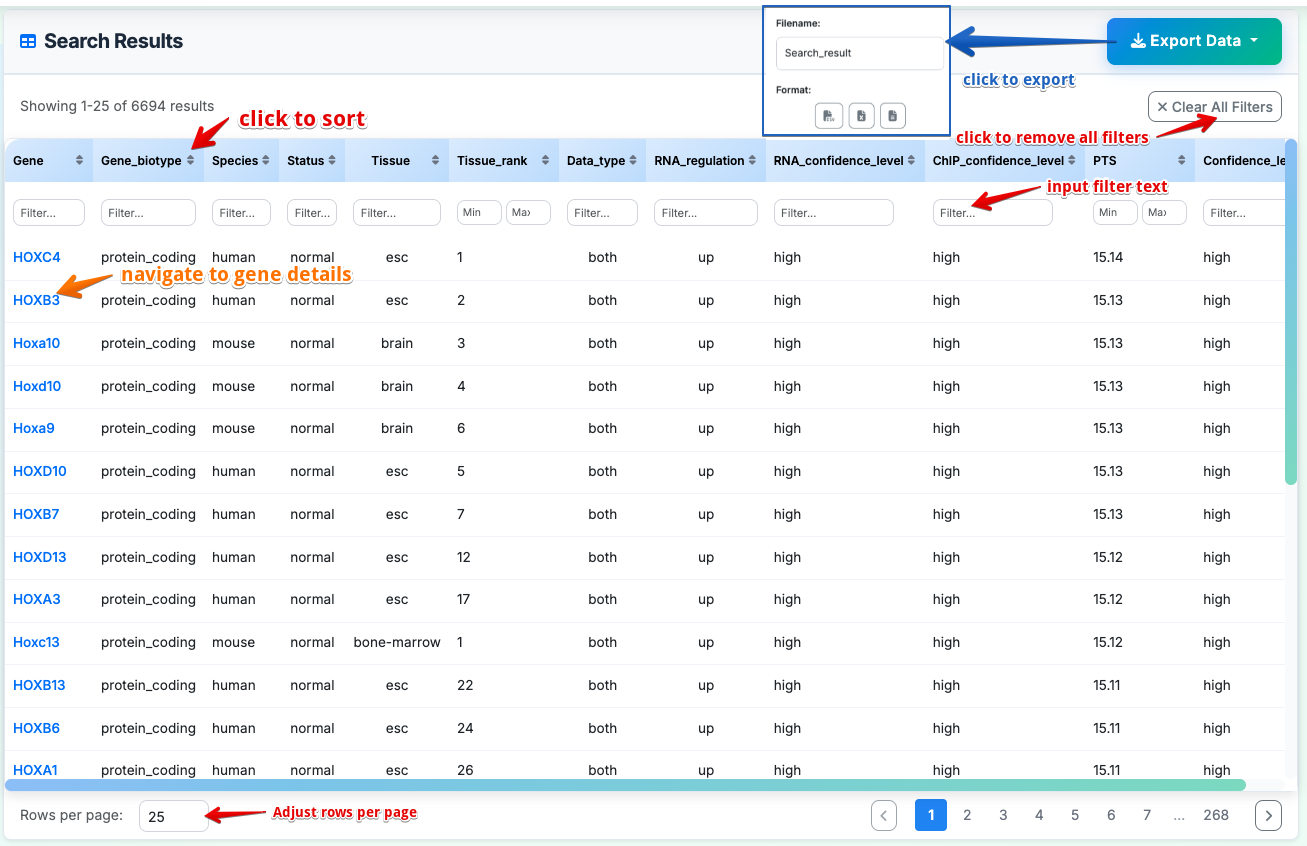
Search Results (Data Page)
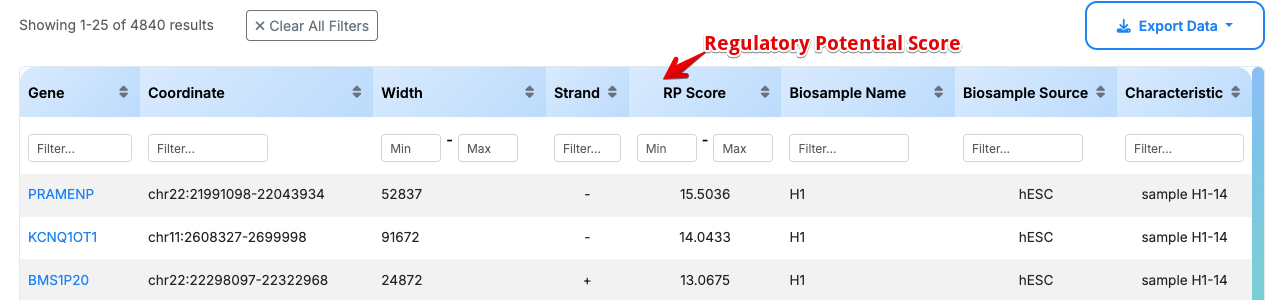
Data Table Columns:
- Gene: Gene symbol (e.g., HOXC4, HOXB3, Hoxa10)
- Gene_biotype: Gene classification (e.g. protein_coding, lncRNA)
- Species: Human or Mouse
- Status: Sample condition (normal/disease)
- Tissue: Tissue type (esc, brain, bone-marrow)
- Tissue_rank: Ranking within tissue
- Data_type: Data source type (RNA-seq, ChIP-seq or both)
- RNA_regulation: Expression direction (up/down)
- RNA_confidence_level: RNA data reliability (high/medium/low)
- ChIP_confidence_level: ChIP data reliability (high/medium/low)
- PTS: Confidence score (e.g., 15, 10)
- Confidence_level: Overall confidence rating (high/medium/low)
Interactive Features:
- Column Filtering: Each column has individual filter options
- Clear All Filters: Reset all applied filters
- Sortable Columns: Click column headers to sort data
- Customizable Display: Adjust rows per page (25 shown)
- Gene Hyperlinks: Click gene symbols to navigate to detailed gene information pages
Search Results (Gene Details Page)
Users can search for genes from the home page or click on any gene name in the results table to access a comprehensive analysis page with seven sections providing detailed gene information and multi-omics data.
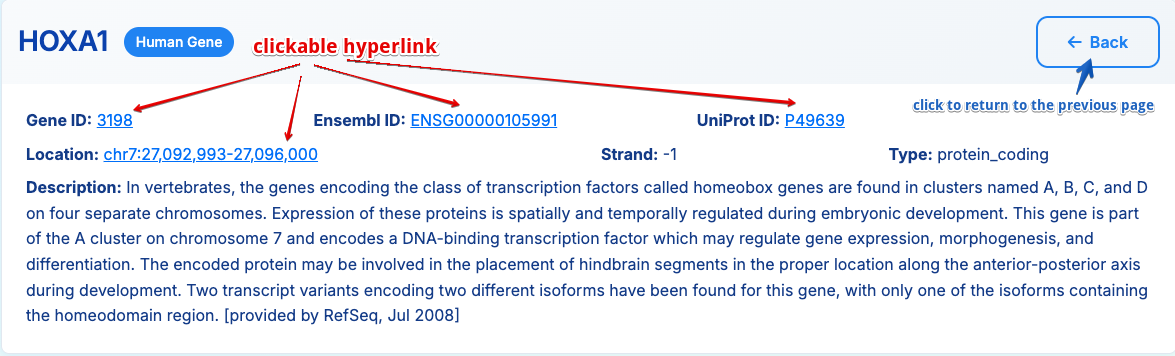
1. Gene Information
Basic gene details including: - Gene symbol, NCBI gene ID, Ensembl ID, UniProt ID and genomic location - External database links (GeneCards, NCBI, Ensembl, UniProt and UCSC DNA sequence)
- Strand orientation and gene type
- NCBI gene functional summary

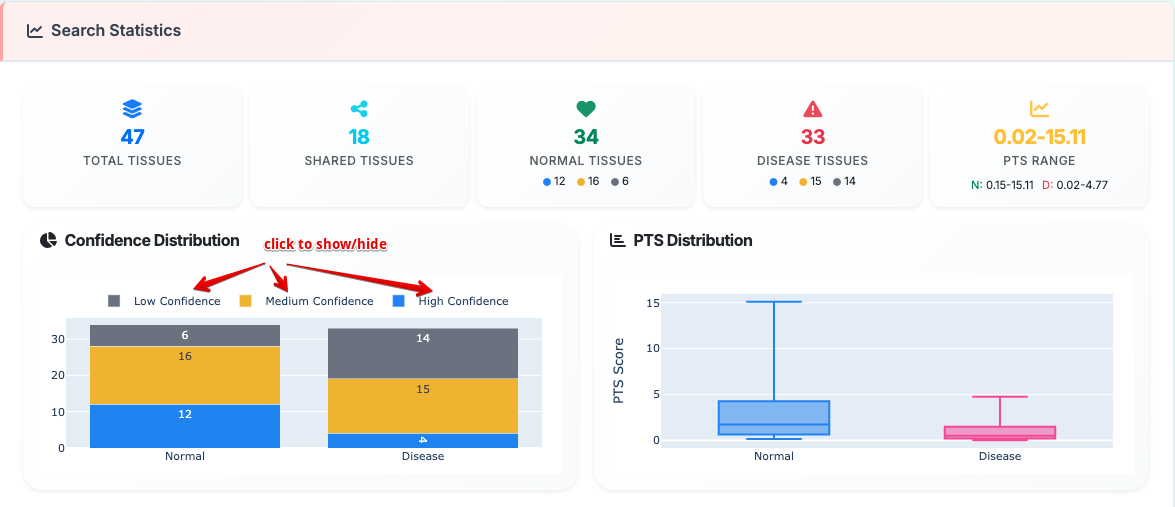
2. Search Statistics
Comprehensive database analytics providing:
- Tissue coverage metrics across normal and disease conditions
- Confidence distribution analysis with quality assessment indicators
- PTS score comparison between sample types
3. PTS Tissue Map
Interactive polycomb target analysis featuring: - Comparative PTS scores across various tissue types - Normal vs Disease condition toggle display - Interactive legend controls for selective data viewing
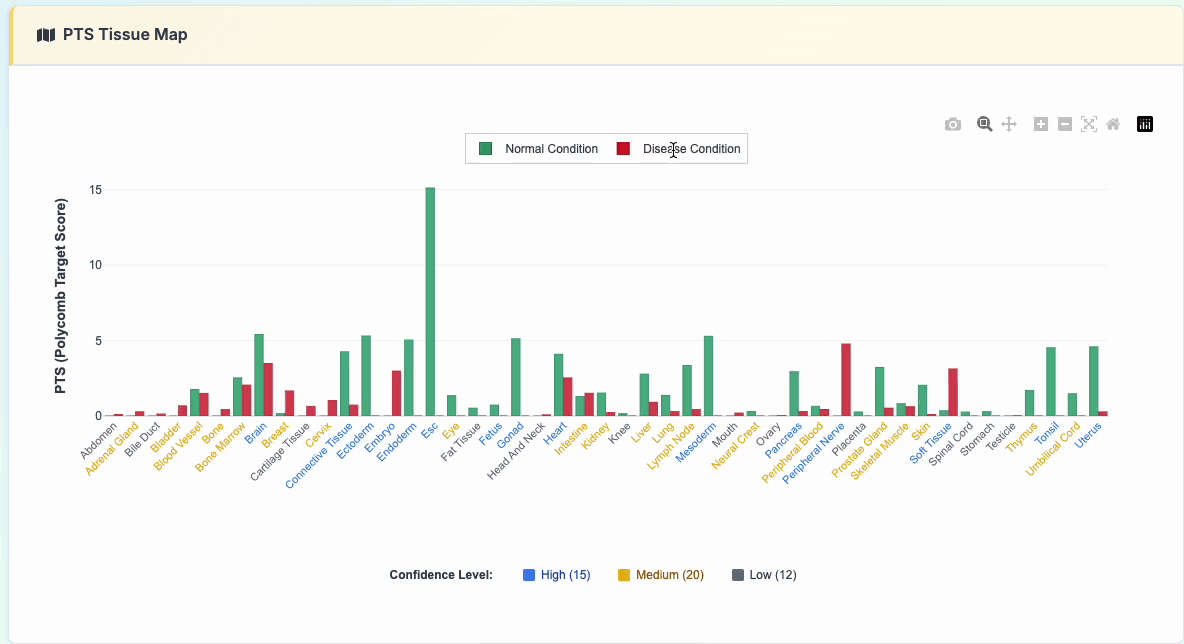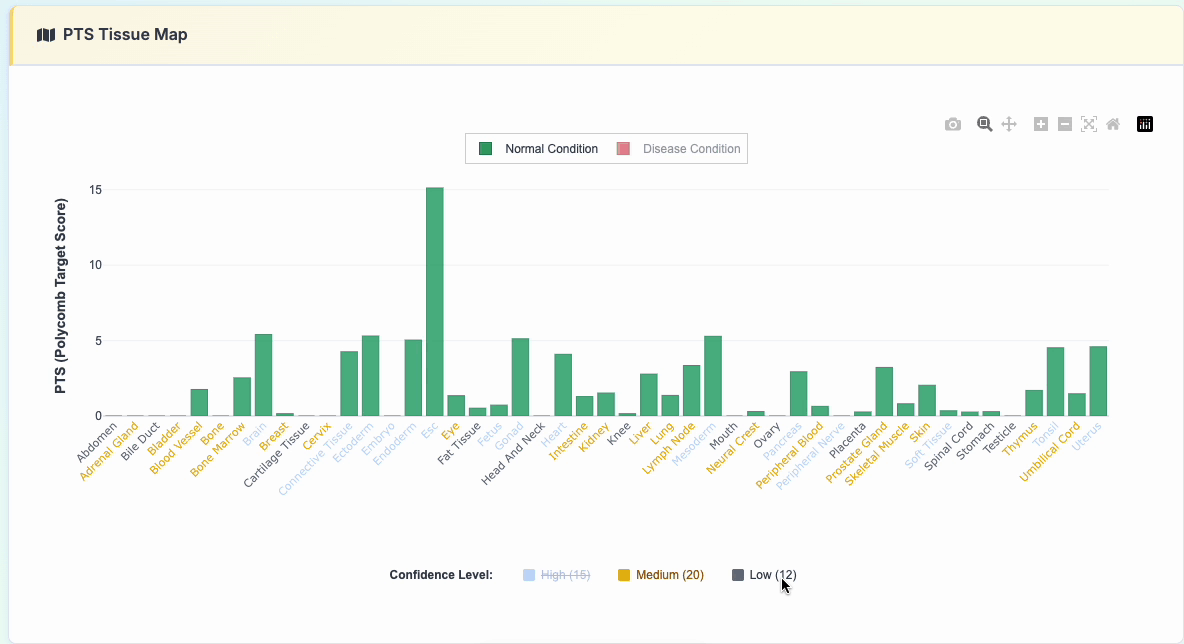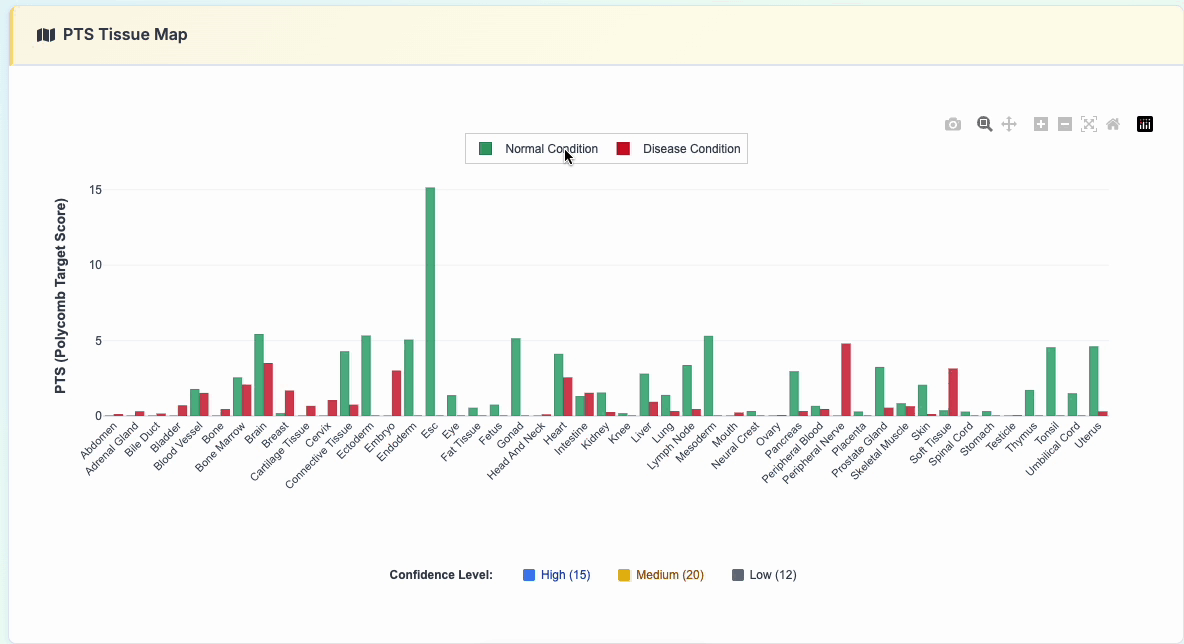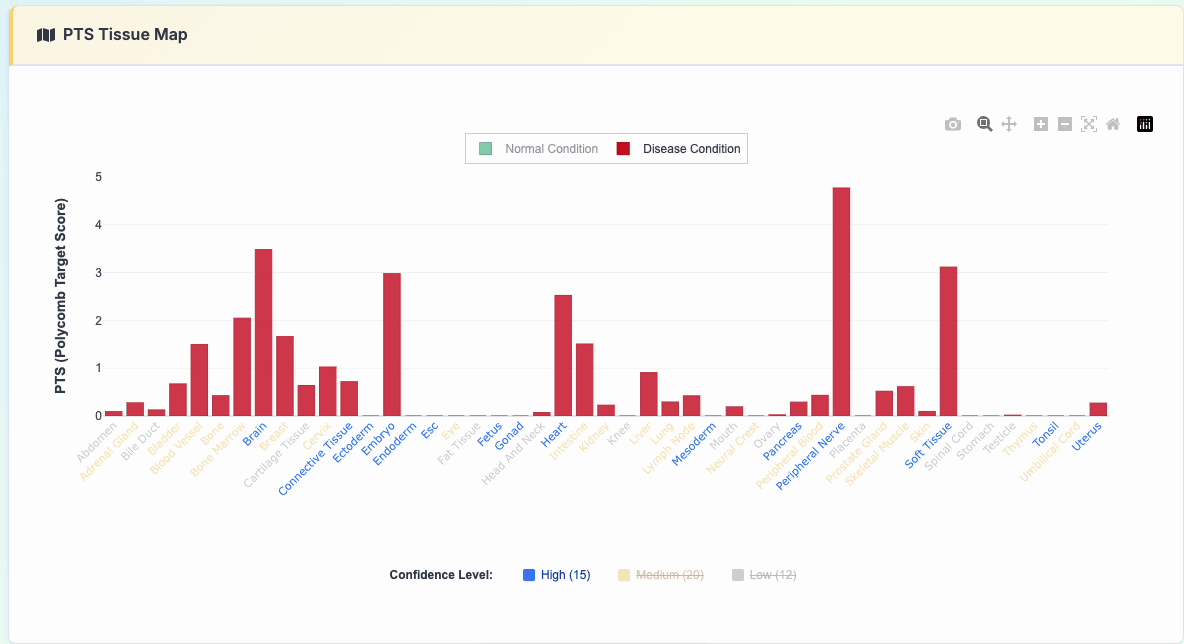
Click the "show details" button to reveal the specific data used:
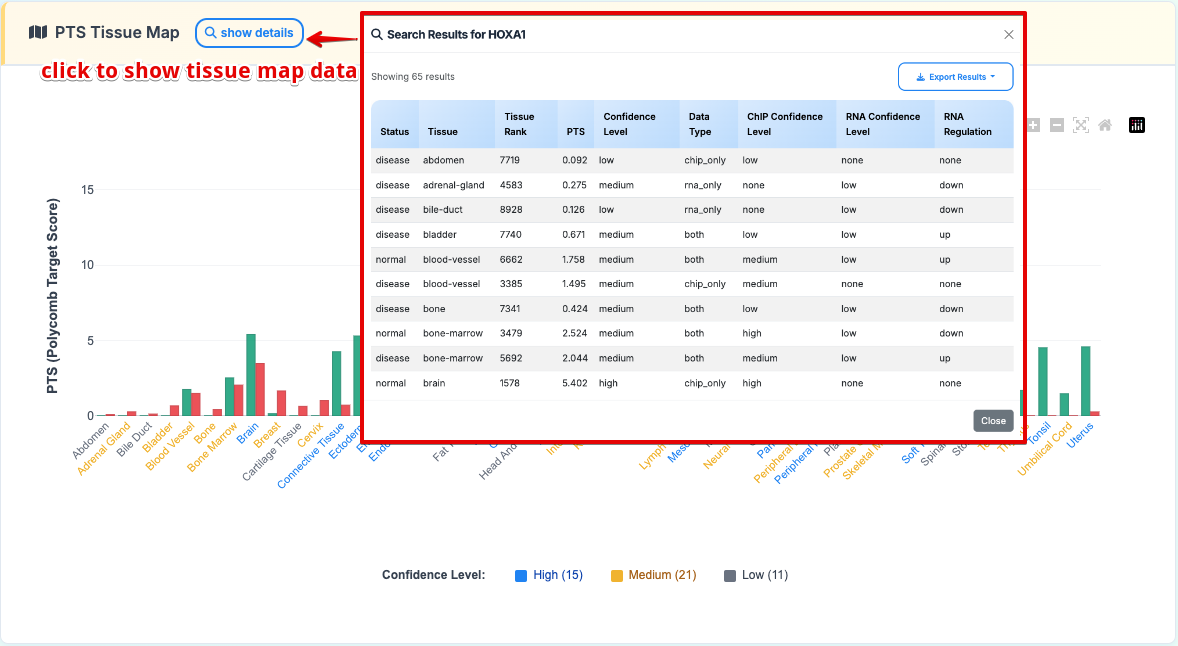
4. Data Details
-
base_id :Unique dataset identifier for traceability
-
biosample_name: Experimental cell/tissue type (e.g., H1-hESC, CD4+ T cells)
-
status: Sample classification (normal vs. disease states)
-
characteristic: Specific experimental conditions or sample annotations
RNA-seq Data Key Columns:
-
treatment: Experimental perturbation targeting Polycomb group proteins
-
mean_exp_control: Baseline expression level in control conditions
-
mean_exp_treat: Expression level following experimental treatment
ChIP-seq Data Key Columns:
- rpscore: Regulatory Potential Score (Higher scores indicate stronger H3K27me3 regulatory potential)
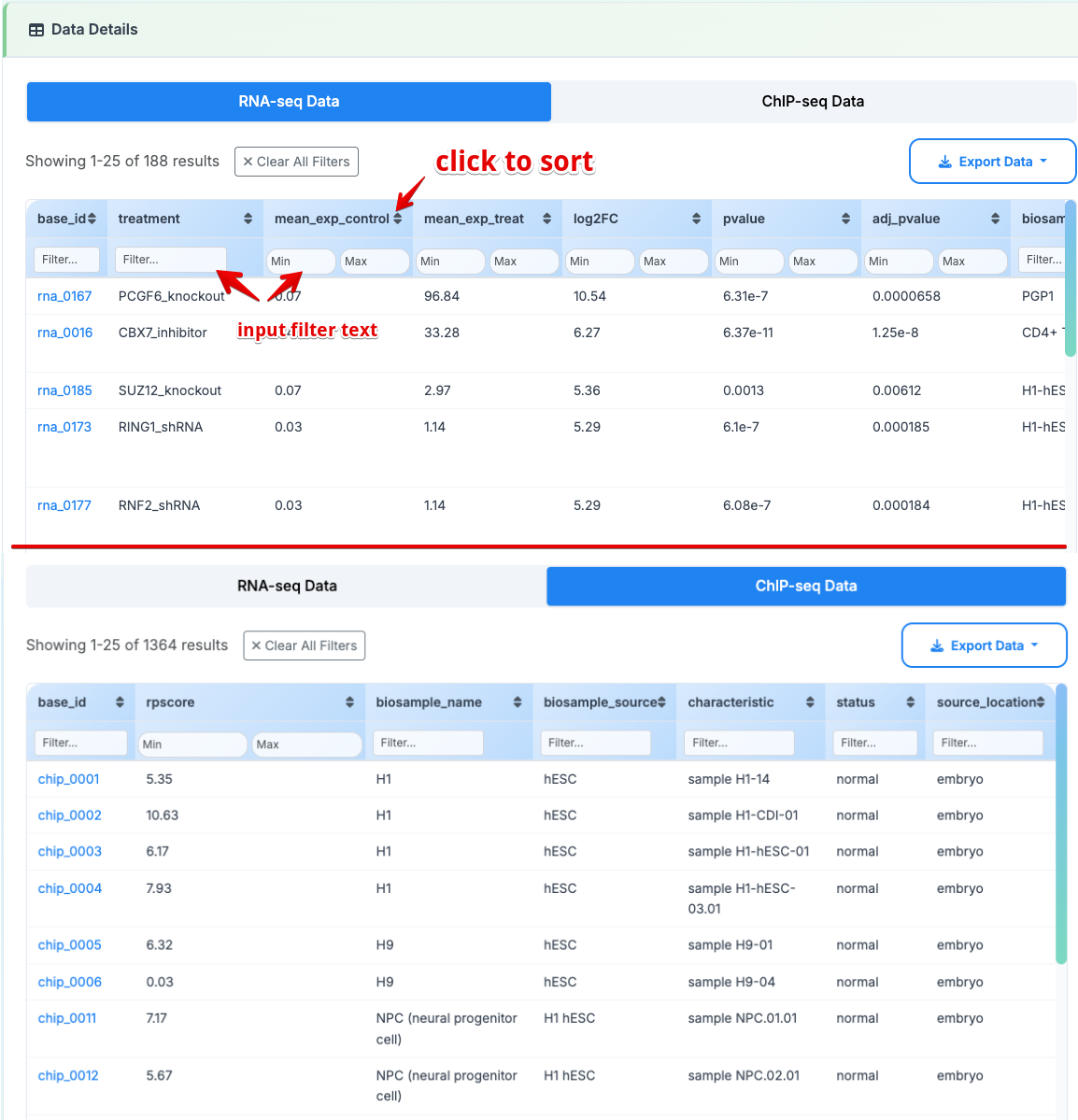
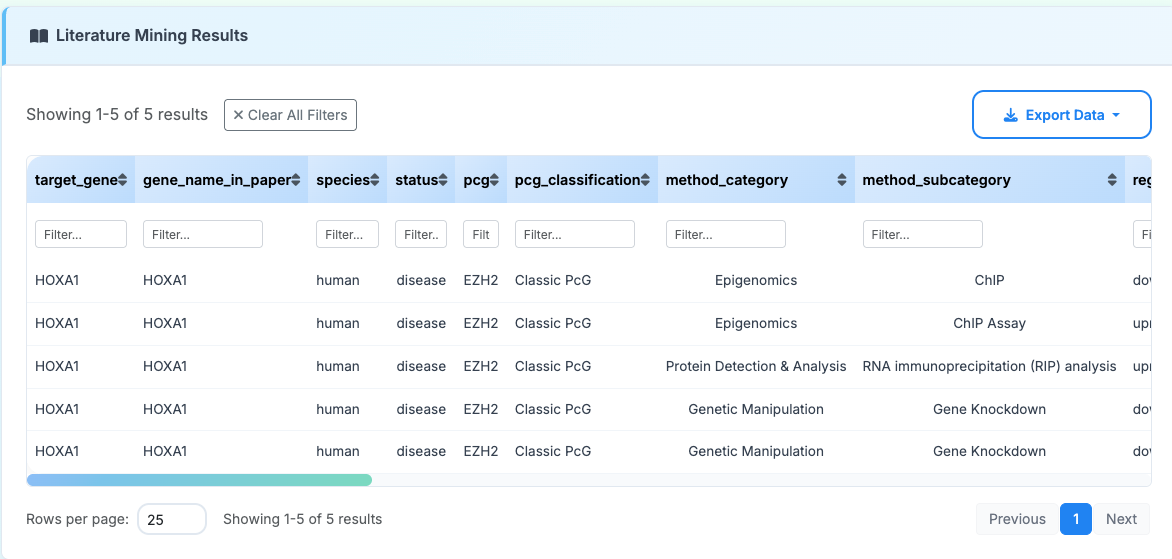
5. Literature Mining
Text mining key columns:
- target_gene: Standardized gene symbol for the target of Polycomb regulation
- Represents genes identified as Polycomb regulatory targets through literature evidence
- gene_name_in_paper: Gene nomenclature as referenced in the original publication
-
May differ from target_gene due to alternative naming conventions or synonyms
-
pcg: Specific Polycomb group protein involved in the regulatory interaction
- Examples: BMI1, SUZ12, EZH2, CBX7
- Identifies which Polycomb component mediates the regulatory effect
- pcg_classification: Functional classification of the Polycomb protein
- Classic PcG:Core components of Polycomb Repressive Complexes with well-established chromatin regulatory functions. [e.g. EZH2, SUZ12, EED, BMI1...]
- Non-classic PcG :Auxiliary regulatory factors or newly discovered Polycomb-associated proteins with emerging functional roles [e.g. RYBP, JARID2, MTF2...]
-
General term:Broad concepts referring to Polycomb complexes, modifications, or collective protein groups rather than specific proteins. [e.g. PRC1, PRC2, PcG, H3K27me3...]
-
method_category: Broad experimental approach used to demonstrate regulation
- Examples: Genetic Manipulation, ChIP-seq, Expression Analysis
- Primary methodology for establishing Polycomb-target relationships
- method_subcategory: Specific experimental technique within the broader category
- Examples: Gene Knockdown, Gene Knockout, Overexpression
- Detailed experimental method for evidence validation
- regulation: Direction of regulatory effect observed
- Examples: upregulated, downregulated, no change
- Indicates whether the target gene’s expression increases or decreases following Polycomb perturbation or direct Polycomb regulation
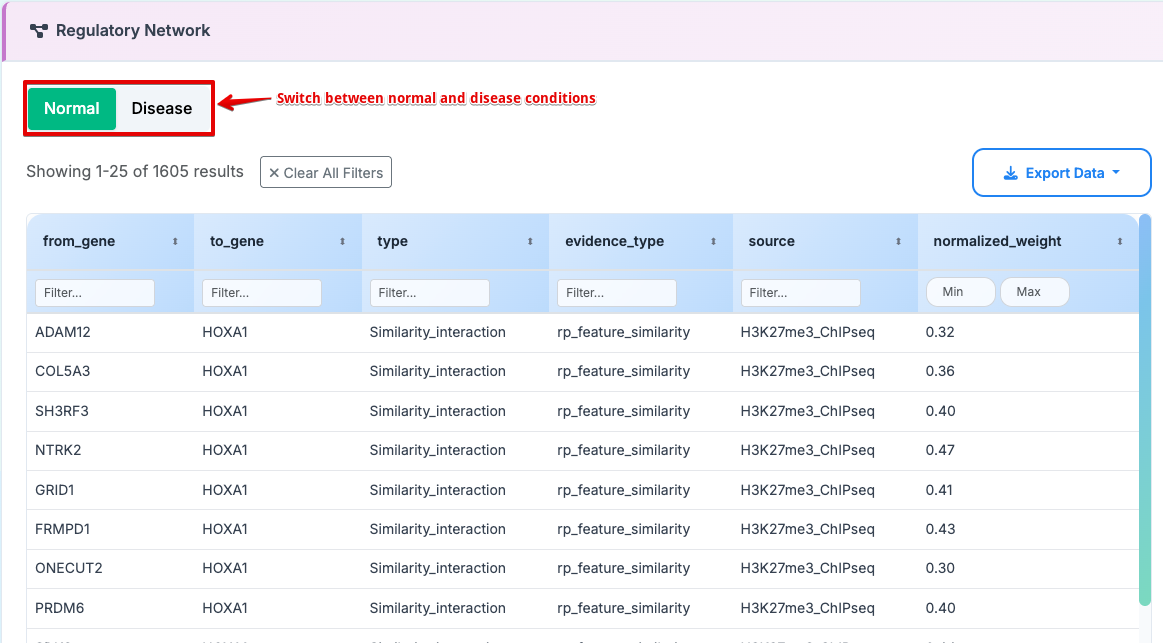
6. Regulatory Network
Gene interaction network columns:
- from_gene: Source gene in the regulatory interaction
- Gene symbol of the upstream regulator or interaction partner
- to_gene: Target gene in the regulatory interaction
-
Gene symbol of the downstream target or interaction partner
-
type: Category of molecular interaction
Similarity_interaction: Co-regulation based on H3K27me3 ChIP-seq similarityPPI_interaction: Protein-protein physical interactionsPcG_regulation: Direct Polycomb group protein regulation-
TF_regulation: Transcription factor regulatory relationships -
evidence_type: Strength and method of experimental support
rp_feature_similarity: Regulatory potential feature similarityprotein_interaction: Physical protein-protein bindinghigh_confidence_regulation: Strong regulatory evidencemedium_confidence_regulation: Moderate regulatory evidencelow_confidence_regulation: Weak regulatory evidencetext_mining: Literature-derived evidence-
molecular_interaction_assay: Direct experimental validation -
source: Database or experimental origin of the interaction
H3K27me3_ChIPseq: Chromatin immunoprecipitation dataSTRING_database: Protein interaction databaseKnock_PcG_RNAseq: Polycomb knockdown RNA-seq experiments-
TRRUST,TFLink,GTRD,ReMap: Transcription factor databases -
weight_normalized: Quantitative strength of the interaction (0-2 scale)
- Higher values indicate stronger regulatory relationships
- Normalized across different evidence types for comparison
- Weight hierarchy:
PcG_regulation > TF_regulation > PPI_interaction > Similarity_interaction
7. Data Visualization
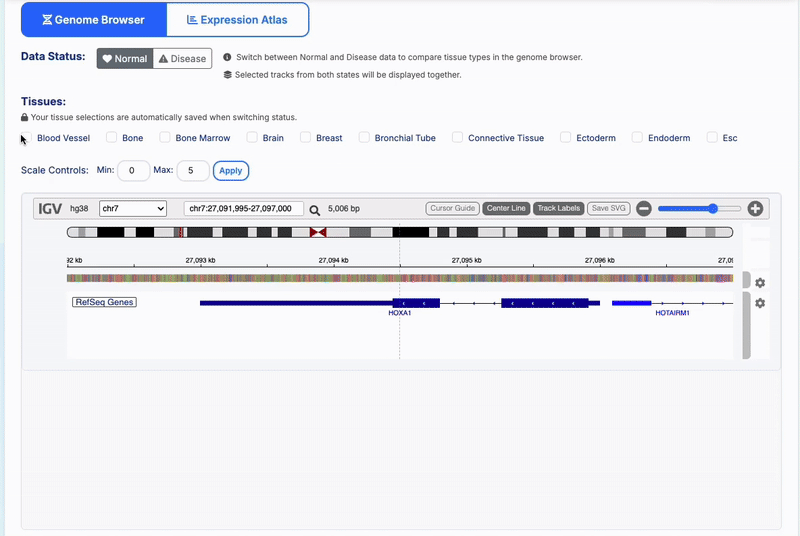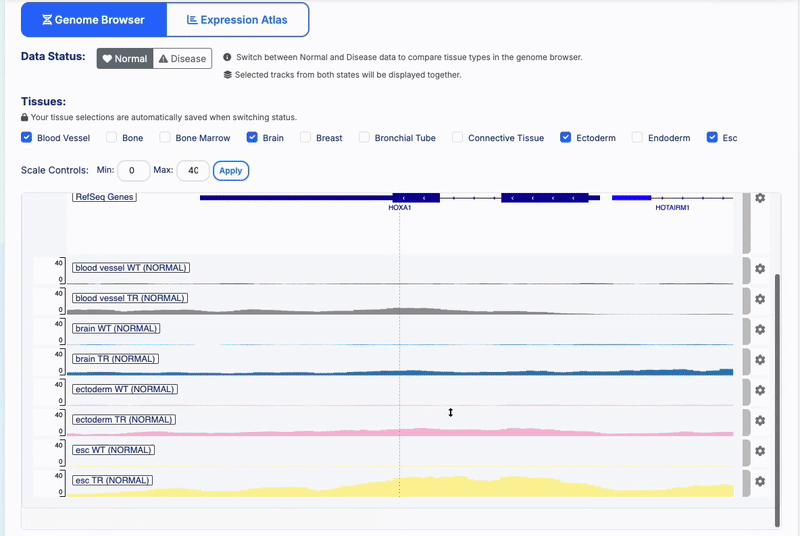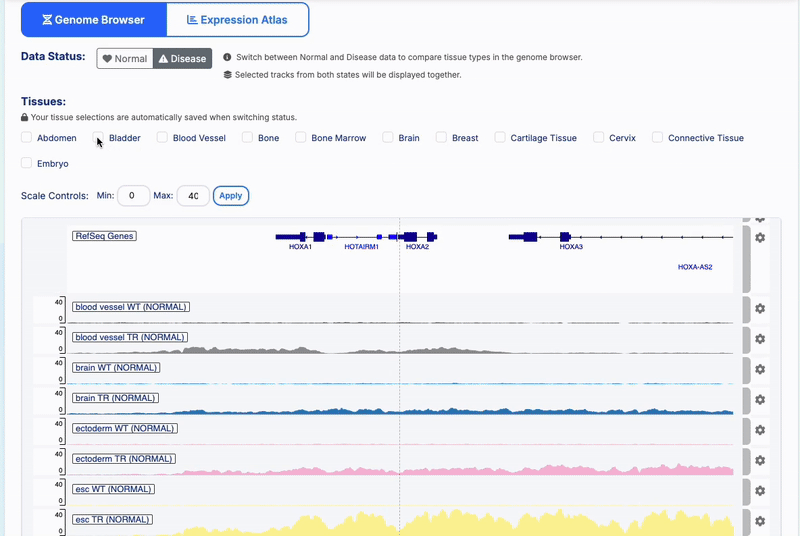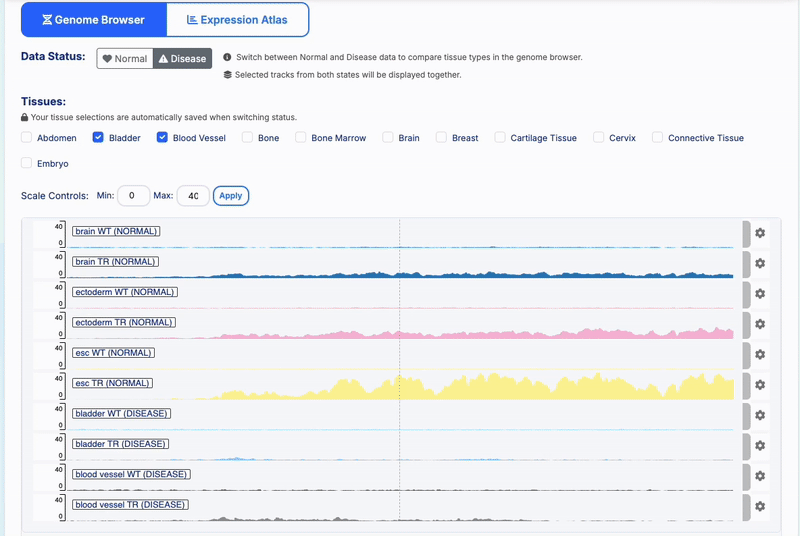
Interactive Genome Browser
- Multi-tissue Support: Select from 30+ human tissues and 20+ mouse tissues
- Dynamic Loading: Tissue options update automatically based on species and data availability
-
State Preservation: Tissue selections are automatically saved when switching between Normal/Disease status
-
Manual Scale Setting: Adjust Min and Max values to normalize all loaded tracks to the same scale range
-
RefSeq Annotations: Gene structure display with exon/intron boundaries
- Interactive Navigation: Search genes, zoom regions, and explore genomic context
- Track Management: Color-coded tissue-specific tracks with WT/TR treatment indicators
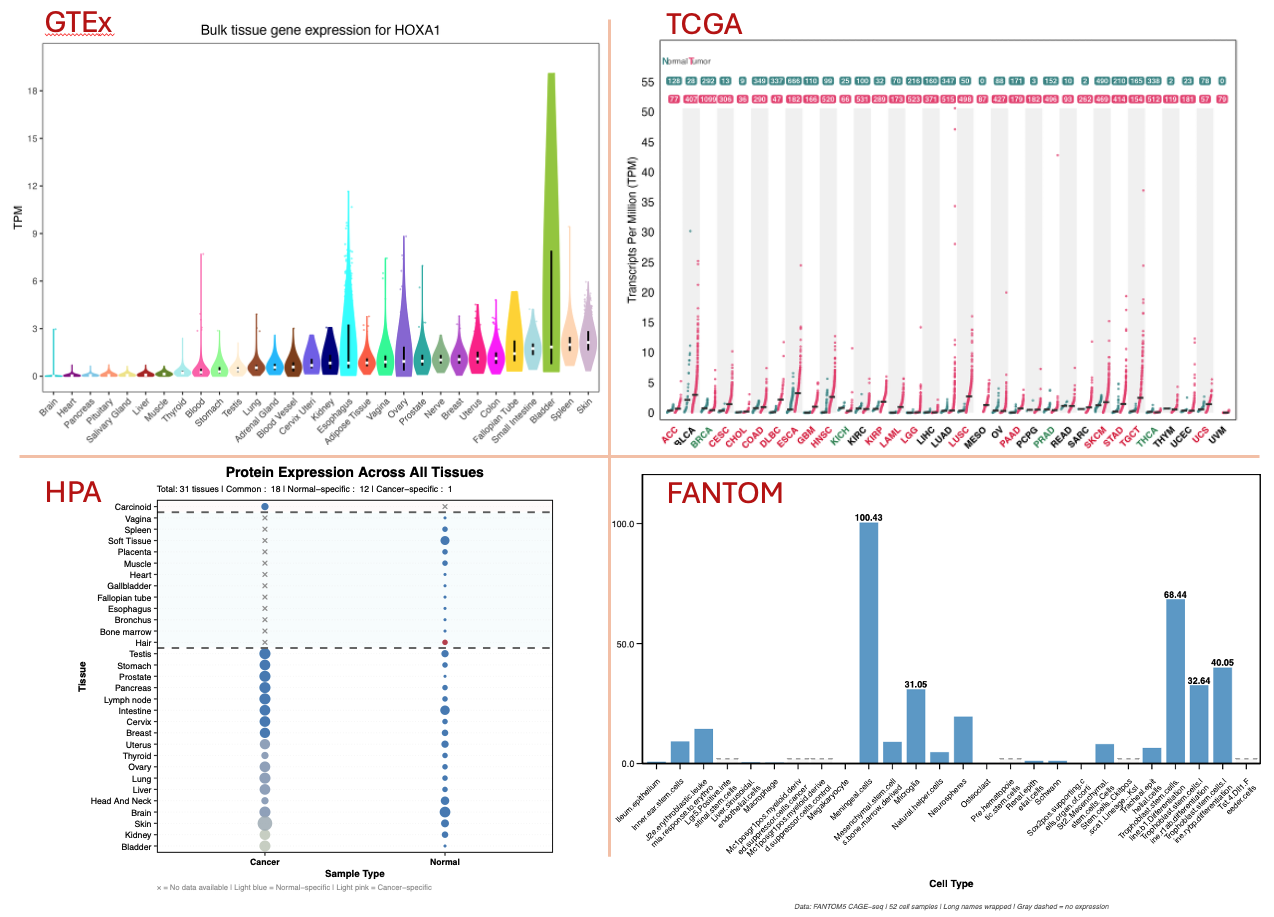
Expression Atlas
[Note: some genes may not have data available in this database, resulting in no image results]
- GTEx (Genotype-Tissue Expression)
- Normal tissue expression across 30+ human tissues
- Violin plots showing expression distribution and median values
-
Tissue-specific color schemes for visual clarity
-
TCGA Pancancer (The Cancer Genome Atlas)
- Cancer tissue expression data across multiple tumor types
- Dot plots comparing normal vs tumor expression levels
-
Disease-specific expression patterns and biomarker identification
-
Human Protein Atlas (HPA)
- Protein-level expression data across human tissues
- Immunohistochemistry and proteomics-based measurements
-
Tissue specificity and subcellular localization information
-
FANTOM5 (Functional Annotation of the Mammalian Genome)
- Mouse tissue and cell-type specific expression
- Two data types:
tissue_levelandcell_levelanalysis - Dual-panel visualization due to extensive tissue coverage
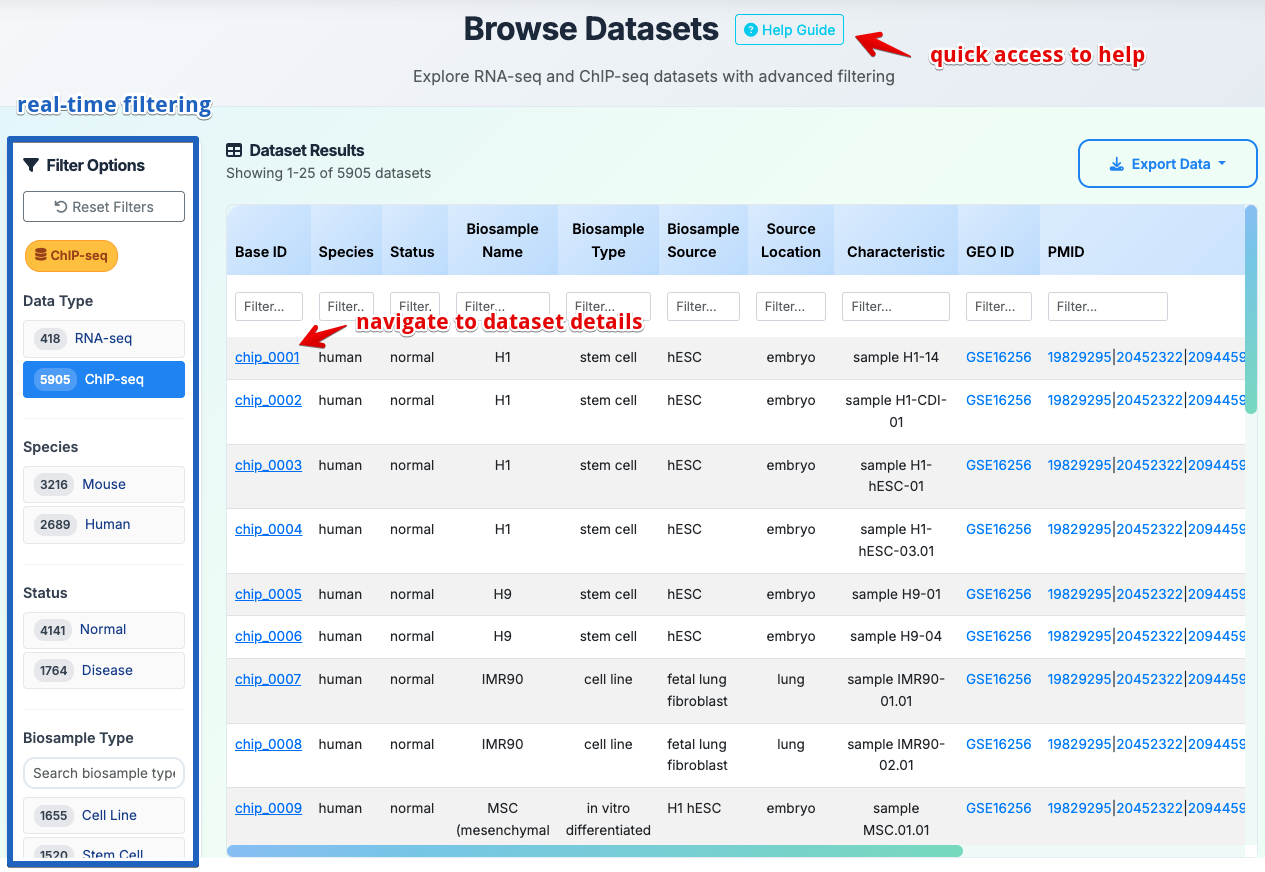
How to Browse Datasets?
Browse datasets of RNA-seq and ChIP-seq experiments with advanced filtering and interactive data exploration.
Filter Options
- Data Type: RNA-seq, ChIP-seq
- Species: Human, Mouse
- Tissue/Cell Type: Organ-specific datasets
- Experimental Conditions: Normal, Disease, Treatment (for RNAseq)
Results Navigation
- Real-time Filtering: Click left panel options, table updates automatically
- Column Sorting: Click column headers for ascending/descending order
- Pagination: Display 10-250 rows per page
- Data Export: CSV, Excel, TXT formats
Browse Results
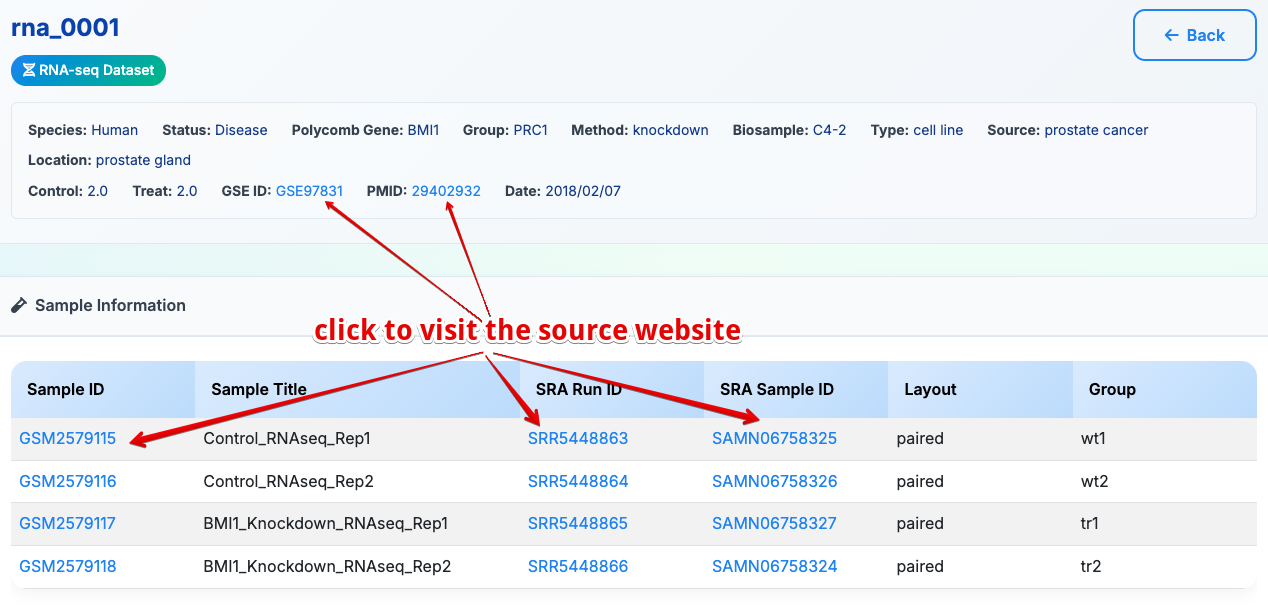
Upon selecting a Dataset ID from the browse results, the user will be directed to a comprehensive dataset analysis page, which is systematically organized into three primary sections:
1. Sample Information
Dataset metadata and experimental details: - Sample ID and external database links - Experimental conditions and treatments - Data type classification (RNA-seq/ChIP-seq) - Publication information and PubMed links
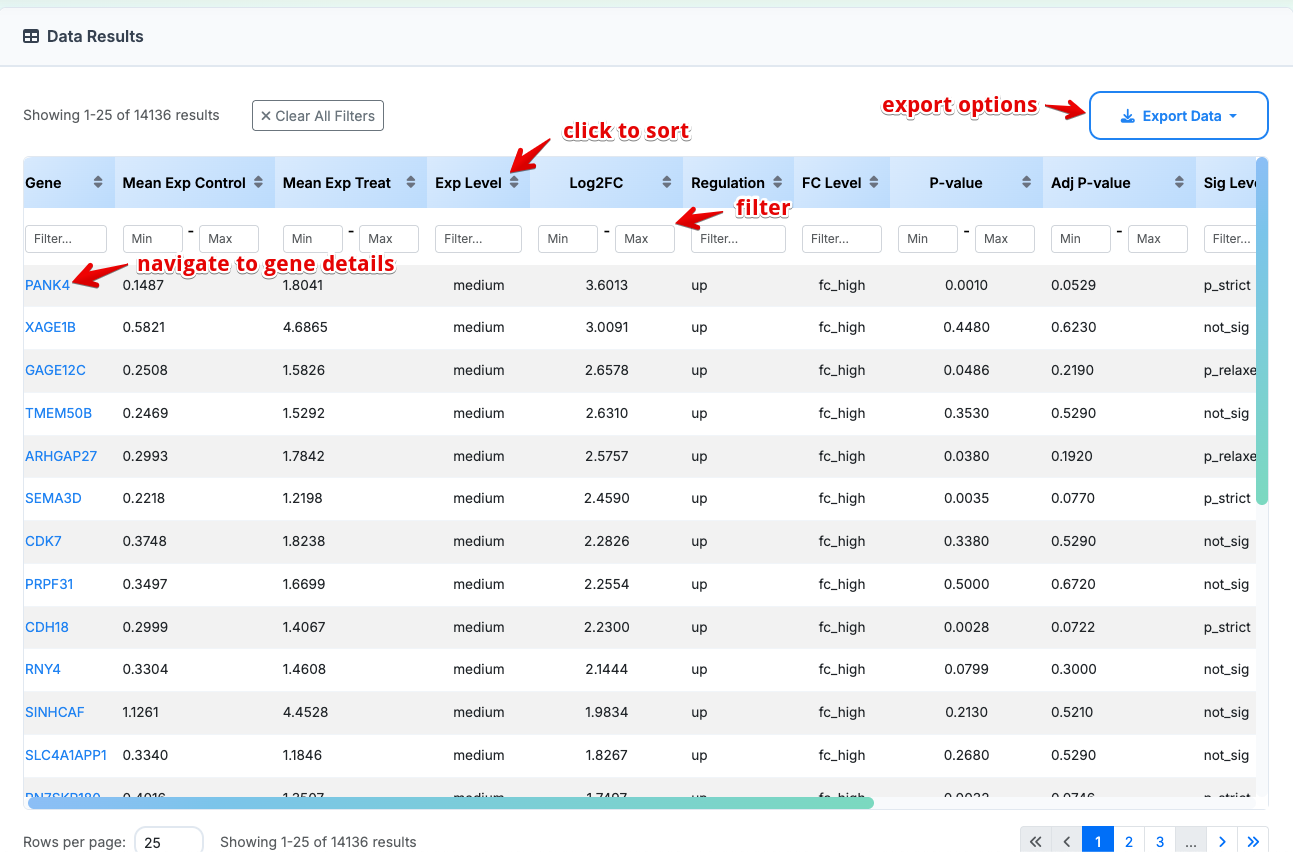
2. Data Details
-
Gene: Gene symbol with a hyperlink to the gene details page
-
Source_location: Tissue or cell type origin
- Status: Disease state (Normal, Disease)
RNA-seq Data Key Columns:
- Mean Exp Control: Mean expression level in control samples
- Mean Exp Treat: Mean expression level in treatment samples
- Exp Level: Expression category (High, Medium, Low)
- High exp: Above 75th percentile in control or treatment group
- Medium exp: Between 35th and 75th percentiles
-
Low exp: Below 35th percentile in control or treatment group
-
FC Level: log2 Fold change magnitude category (High FC, Medium FC, Low FC, No change)
- High FC: Above 75th percentile [Strong biological relevance]
- Medium FC: Above 50th percentile [e.g.
log2FC ≈ 0.585 equals 1.5-fold change] - Low FC: Above 25th percentile [e.g.
log2FC ≈ 0.265 equals 1.2-fold change] -
No Change: Below all dynamic thresholds [Changes likely from technical noise]
-
Sig Level: Statistical Significance Levels
- FDR Strict (fdr_strict):
adj_pvalue < 0.01 - FDR Relaxed (fdr_relaxed):
adj_pvalue < 0.05 - P-value Strict (p_strict):
pvalue < 0.01 (when FDR unavailable) - P-value Relaxed (p_relaxed):
pvalue < 0.05 (when FDR unavailable) - Not Significant (not_sig): Insufficient statistical evidence
- Confidence Level: Quickly assess data quality
- High Confidence:
FDR < 0.01 AND High fold change AND High expression - Medium Confidence:
Strict significance (FDR/p < 0.01) AND Medium fold changeorStandard significance (FDR/p < 0.05) AND Medium/High fold change - Low Confidence:
Any significance level AND Low fold changeorWeak significance AND Medium fold changeorObservable trend without statistical support
ChIP-seq Data Key Columns:
RP Score (Regulatory Potential Score):Strength of protein binding at genomic regions detected by ChIP-seq experiments. Higher scores = stronger binding.
RP_Score = Σ(distance_score × peak_signal) / √(number_of_peaks)
- distance_score: Location-based weight
- peak_signal: ChIP-seq signal intensity at peak
- √(number_of_peaks): Normalization to prevent long genes from getting artificially high scores
Distance Score Calculation:
- Promoter Region (Direct transcription control): distance_score =
exp(-0.004 × |distance_to_TSS|) - Exponential decay with distance; Closer peaks = higher score
-
Example:
TSS (distance=0) → score=1.0;1000bp away → score=0.98 -
Intragenic Region: (Transcription elongation/splicing ) distance_score =
0.4 (fixed weight) -
Moderate regulatory potential; Affects transcription elongation/splicing
-
Intergenic Region (Distant regulatory elements): distance_score =
0.3 (fixed weight) - Lower regulatory potential; Distant enhancers or weak effects
Example: For a gene with 3 H3K27me3 peaks:
Peak 1: Promoter, distance=500bp, signal=100
→ distance_score = exp(-0.004 × 500) = 0.135
→ contribution = 0.135 × 100 = 13.5
Peak 2: Intragenic, signal=80
→ distance_score = 0.4
→ contribution = 0.4 × 80 = 32.0
Peak 3: Intergenic, signal=60
→ distance_score = 0.3
→ contribution = 0.3 × 60 = 18.0
RP_Score = (13.5 + 32.0 + 18.0) / √3 = 63.5 / 1.73 = 36.7
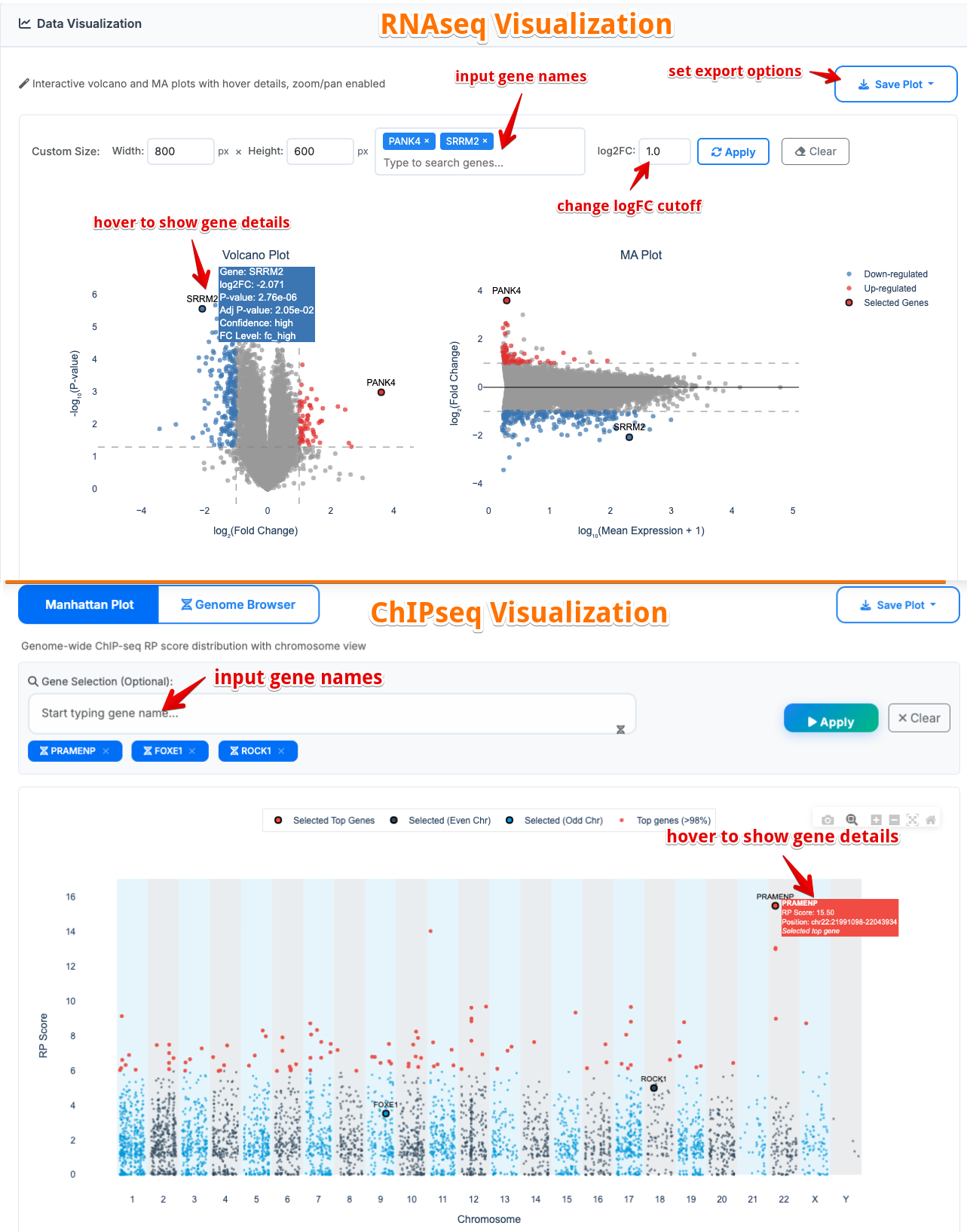
3. Data Visualization
Interactive plots showing dataset characteristics:
- Volcano Plot: Differential expression analysis
- MA Plot: Mean expression vs fold change
- Manhattan Plot: Genome-wide significance mapping
- Custom plot dimensions and export options (PNG/PDF/SVG/HTML)
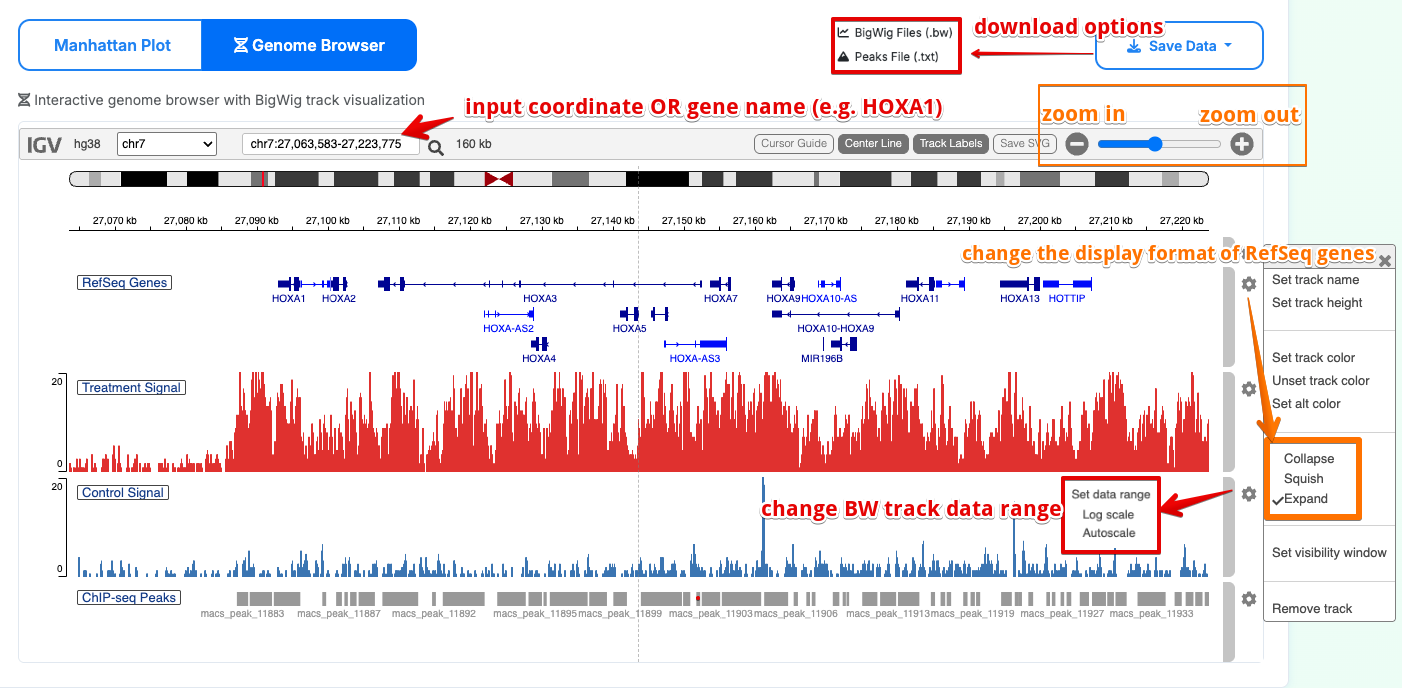
Interactive genomic data visualization:
- Automatically load hg38 or mm10 according to the species of the ChIP-seq dataset
- Scale normalization and track management
- Real-time genomic region navigation
- Provide complete downloads for BW and peak files
How to Analyze Data?
PRIME provides four analysis tools to explore Polycomb targets. Click on each module below for detailed instructions.

Comparison Analysis
Compare targets across different conditions and tissues to identify various polycomb regulatory patterns.
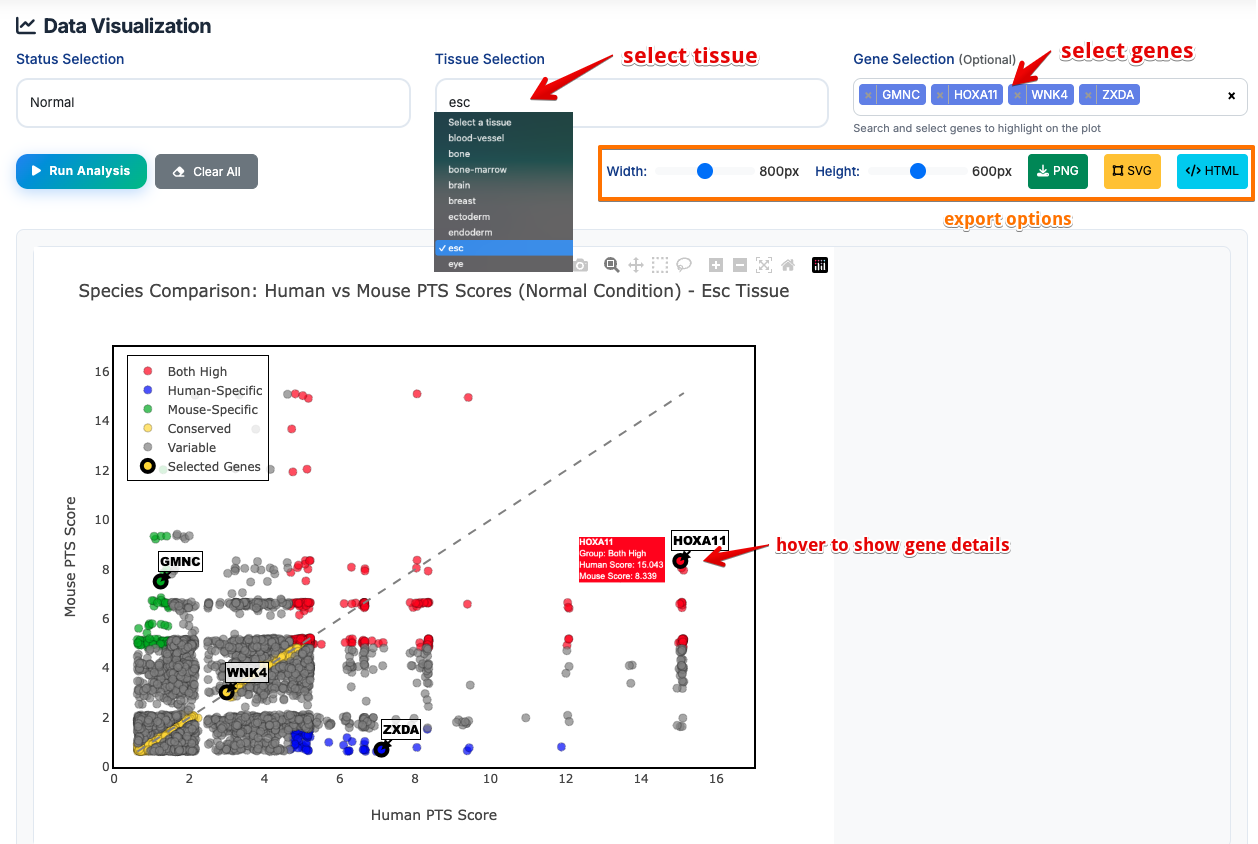
Sub-Module 1: Species Comparison
Purpose: compare the same tissue (human vs mouse) and find cross-species conserved targets
Usage:
- Select status: Choose normal or disease from the dropdown menu
- Choose tissue: Select one tissue
- Input gene name (optional): Input one or more genes to display on the plot
- Run Analysis: Click the button to generate dynamic plots
- Download Results: Export volcano plots and MA plots in PNG/SVG/HTML formats
- Clear All: Reset all parameters and clear the image
Scatter plot result:
- Both High (Red Points):
human_score > 80th percentileANDmouse_score > 80th percentile - Meaning: Genes with strong Polycomb regulation in both species
- Location: Upper right quadrant
- Human-Specific (Blue Points):
human_score > 80th percentileANDmouse_score < 20th percentile - Meaning: Strong Polycomb regulation only in human
- Location: Lower right quadrant
- Mouse-Specific (Green Points):
human_score < 20th percentileANDmouse_score > 80th percentile - Meaning: Strong Polycomb regulation only in mouse
- Location: Upper left quadrant
- **Conserved (Yellow Points): **
|human_score - mouse_score| < 25th percentile of all differences - Meaning: Similar expression levels between species (regardless of absolute level)
- Location: Along diagonal line
- **Variable (Gray Points): ** All other genes not fitting above categories
Sub-Module 2: Tissue Comparison
Purpose: compare Polycomb Target Scores (PTS) across different tissues within normal and disease status to identify tissue-specific Polycomb regulatory patterns.
Usage:
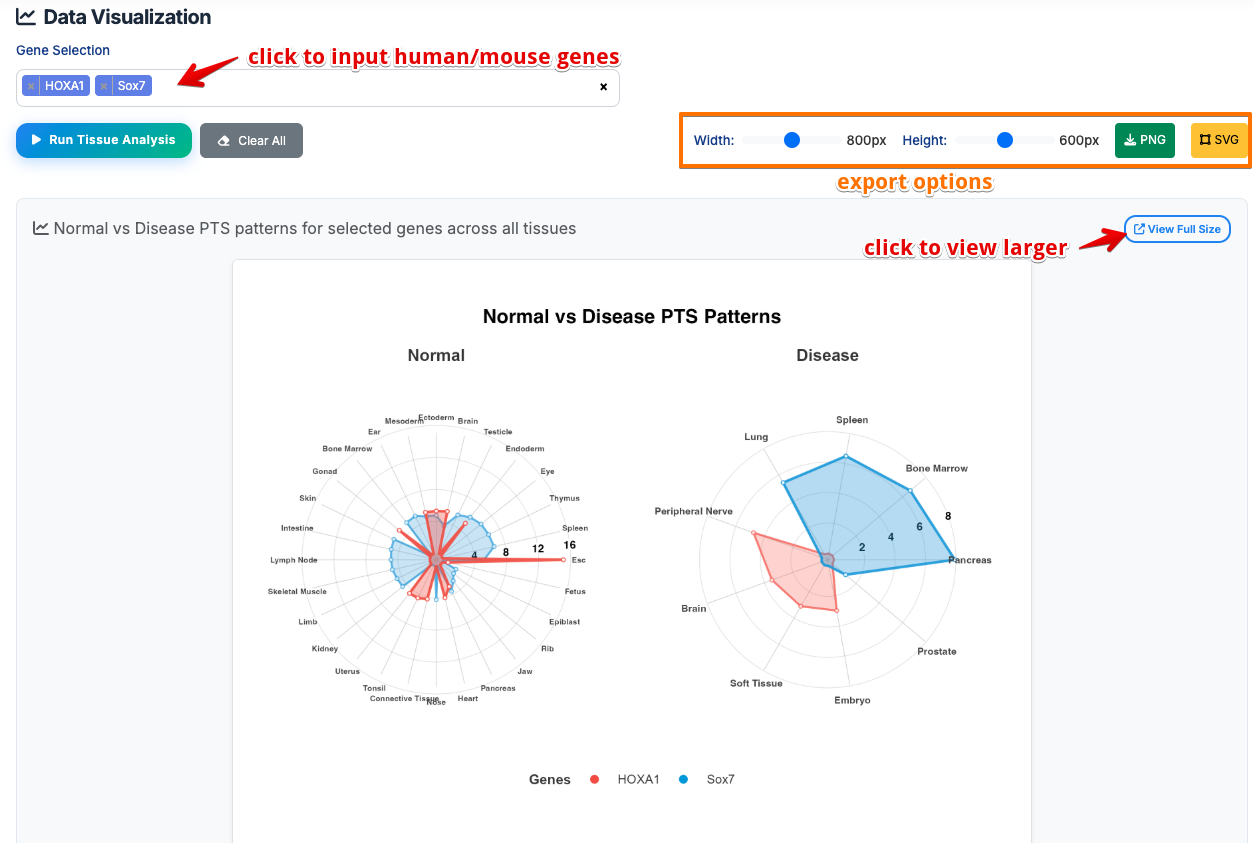
- Input Gene Names: Enter one or more human/mouse gene symbols (e.g., HOXA1, Sox7)
- Run Analysis: Click button to generate interactive radar charts
- Download Results: Export radar plots in PNG/SVG formats
Radar Chart Results:
- Tissue-Specific Targeting: Sharp spikes in specific directions
-
Meaning: Strong Polycomb regulation limited to few tissues
-
Broad Polycomb Targeting: Circular/symmetric polygon shape
- Meaning: Gene is consistently targeted by Polycomb across multiple tissues
Sub-Module 3: Status Comparison
Purpose: compare Polycomb Target Scores (PTS) between Normal and Disease conditions within tissues to identify disease-associated changes in Polycomb regulation.
Usage:
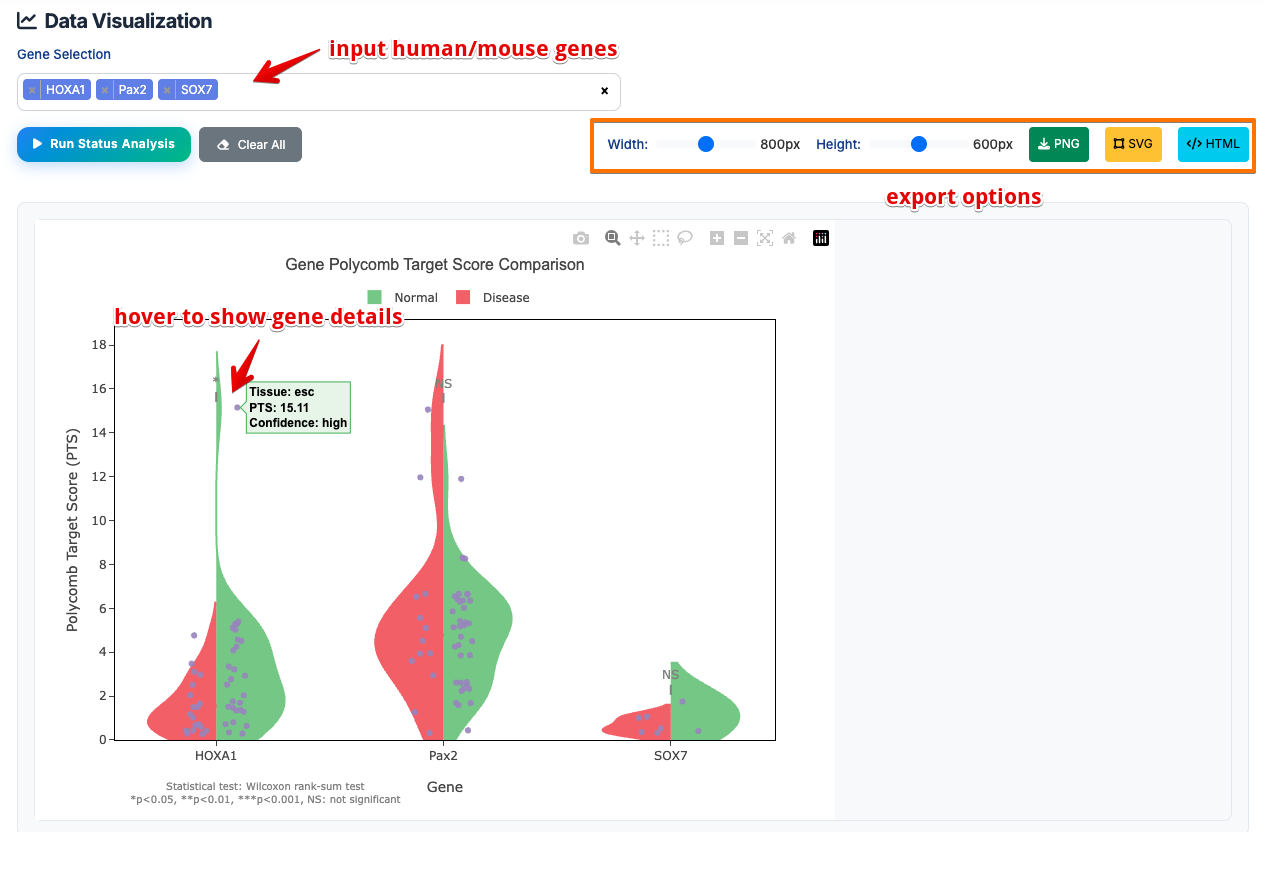
- Input Gene Names: Enter one or more human/mouse gene symbols (e.g., HOXA1, Pax2, SOX7)
- Run Analysis: Click button to generate interactive violin plots
- Download Results: Export violin plots in PNG/SVG/HTML formats
Violin Plot Results:
- Normal Status (Right Side/Green): Shows PTS distribution in healthy conditions
- Disease Status (Left Side/Red): Shows PTS distribution in disease conditions
- Each point represents the PTS value of a tissue, and users can hover to view the details.
Network Analysis
Purpose: Build regulatory networks to view hub genes, analyze protein-protein interactions, and explore Polycomb-mediated regulatory relationships within selected gene sets.
Usage:
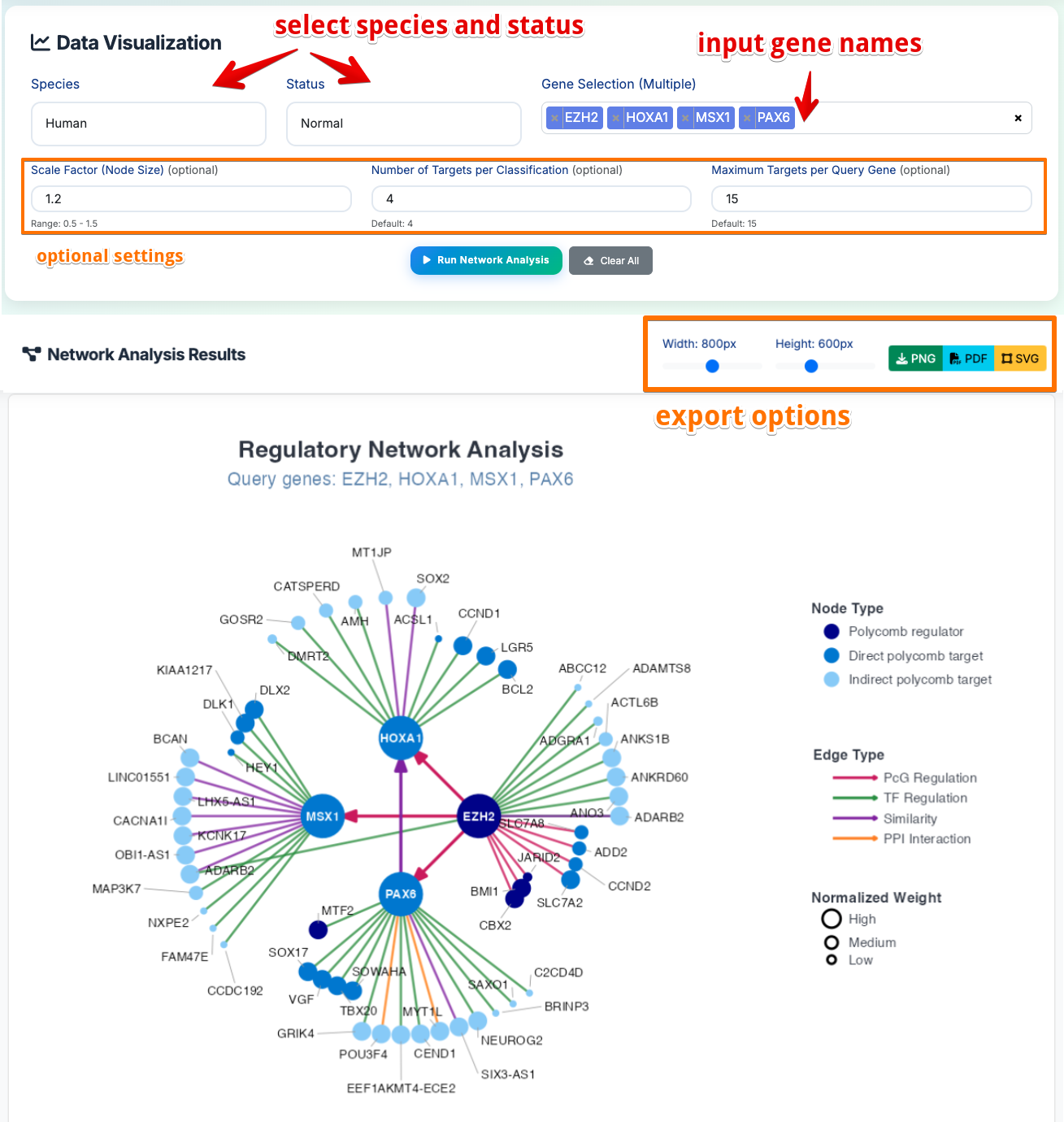
- Select Species: Choose Human or Mouse from dropdown menu
- Select Status: Choose Normal or Disease condition
- Input Query Genes: Enter gene symbols of interest (e.g., EZH2, MSX1, HOXA1, PAX6)
- Set Parameters: Adjust hub gene degree threshold and maximum targets per query
- Run Analysis: Click button to generate interactive network diagram
- Download Results: Export network plots in PNG/PDF/SVG formats
Network Visualization Results:
- Central Hub Genes (Large Dark Nodes): Highly connected regulatory genes
- Meaning: Master regulators with extensive downstream networks
-
Location: Central positions with many outgoing connections (e.g., PcG components such as EZH2)
-
Query Genes (Large Colored Nodes): User-specified genes of interest
- Meaning: Starting points for network exploration
-
Location: Prominently sized nodes with gene labels (HOXA1, MSX1, PAX6)
-
Target Genes (Small Nodes): Downstream regulated genes
- Meaning: Genes regulated by hub/query genes in the network
-
Location: Smaller peripheral nodes connected via edges
-
Node Colors by Function:
- Dark Blue: Polycomb regulators (master regulatory genes)
- Medium Blue: Direct Polycomb targets
- Light Blue: Indirect Polycomb targets
- Edge Colors by Interaction Type:
- Red: Polycomb-mediated regulation (PcG_regulation)
- Green: Transcription factor regulation (TF_regulation)
- Purple: Expression similarity patterns (Similarity_interaction)
-
Orange: Protein-protein interactions (PPI_interaction)
-
Node Size: Represents interaction confidence
- Circle size indicates normalized weight (High/Medium/Low)
Disease & Drug Analysis
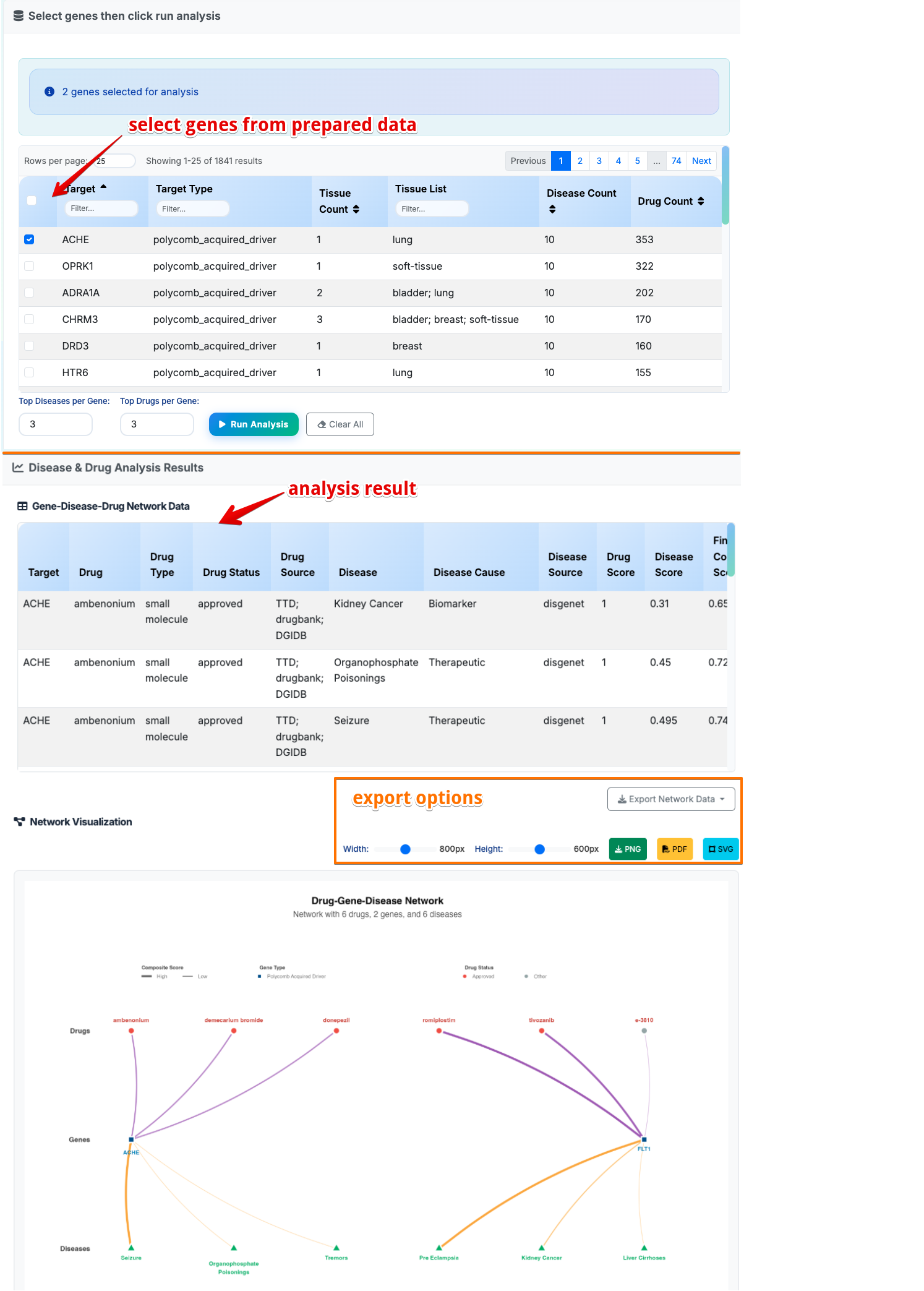
Purpose: Analyze the relationships between polycomb target genes, associated diseases, and candidate drugs. The module uses computational databases to build gene-disease-drug networks and prioritize research opportunities based on composite scoring systems.
Usage:
- Browse Gene Targets: The interface loads 1,841 polycomb target genes with disease associations
- Target Type:
- Polycomb-acquired drivers are genes that newly gain Polycomb-mediated silencing in disease, often acting as causal factors;
- Polycomb-perturbed biomarkers are genes whose pre-existing Polycomb regulation becomes significantly altered in disease, serving as markers of disease response.
- Tissue Count: Number of tissues where the target is active
- Tissue List: Specific tissues showing polycomb regulation
- Disease Count: Number of associated diseases
- Drug Count: Number of potential therapeutic compounds
- Select Genes: Use table filtering and checkboxes to select genes of interest
- Set Parameters: Adjust top diseases per gene (default: 3) and top drugs per gene (default: 3)
- Run Analysis: Click button to generate comprehensive results
Gene-Disease-Drug Network Data Results:
- Target: Selected polycomb target gene
- Drug: Candidate therapeutic compound
- Drug Type: Classification of drug mechanism
- Drug Status: Development stage (e.g. approved, investigational, experimental)
- Drug Source: Database source of drug information (TTD, DrugBank, DGIDB)
- Disease: Associated disease name
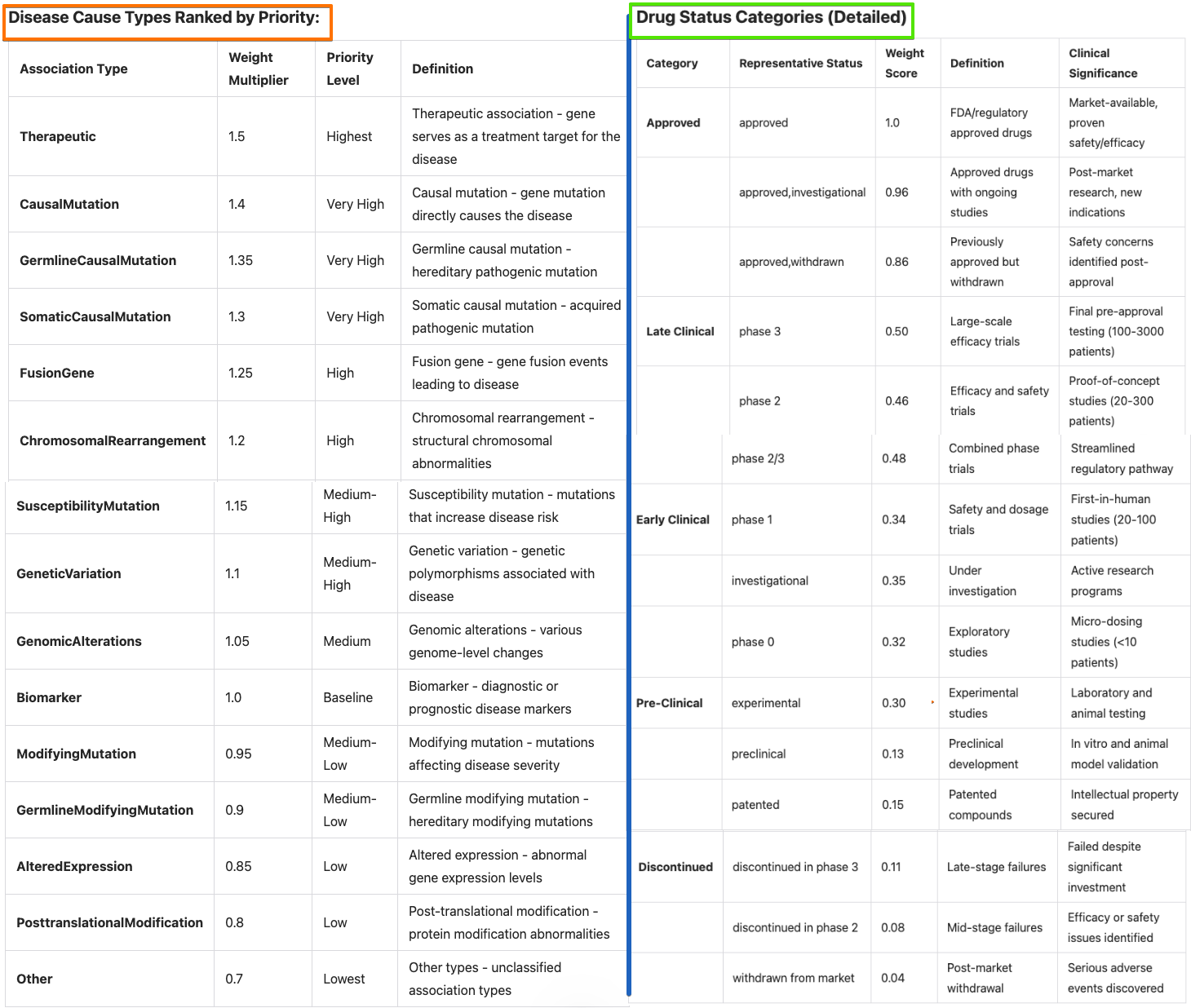
- Disease Cause: Pathological mechanism category (e.g. Mutation, Biomarker)
- Disease Source: Database source of disease association (DisGeNET, Orphanet)
- Drug Score: Computational drug-target interaction score
- Disease Score: Gene-disease association strength score
- Final Composite Score: Integrated prioritization score
- Research Priority: Overall ranking for therapeutic development (
Final Composite Score >= 0.7 ~ High Priority;Final Composite Score >= 0.4 ~ Medium Priority) - Target Type: Polycomb classification category
Gene-Disease-Drug Networ Visualization:
- Drug Nodes (Circle): Therapeutic compounds targeting query genes
- Meaning: Potential drugs for treating Polycomb-related diseases
- Central Gene Nodes (Square): User-selected query genes of interest
- Meaning: Polycomb targets with disease/drug associations
- Disease Nodes (Triangle): Disease conditions associated with query genes
- Meaning: Pathological conditions linked to Polycomb dysregulation
Motif Analysis
Identify transcription factor binding sites within polycomb target gene promoter, providing insights into regulatory mechanisms. The module integrates JASPAR motif databases to analyze gene-motif relationships across human and mouse species, supporting both individual gene analysis and comparative studies.
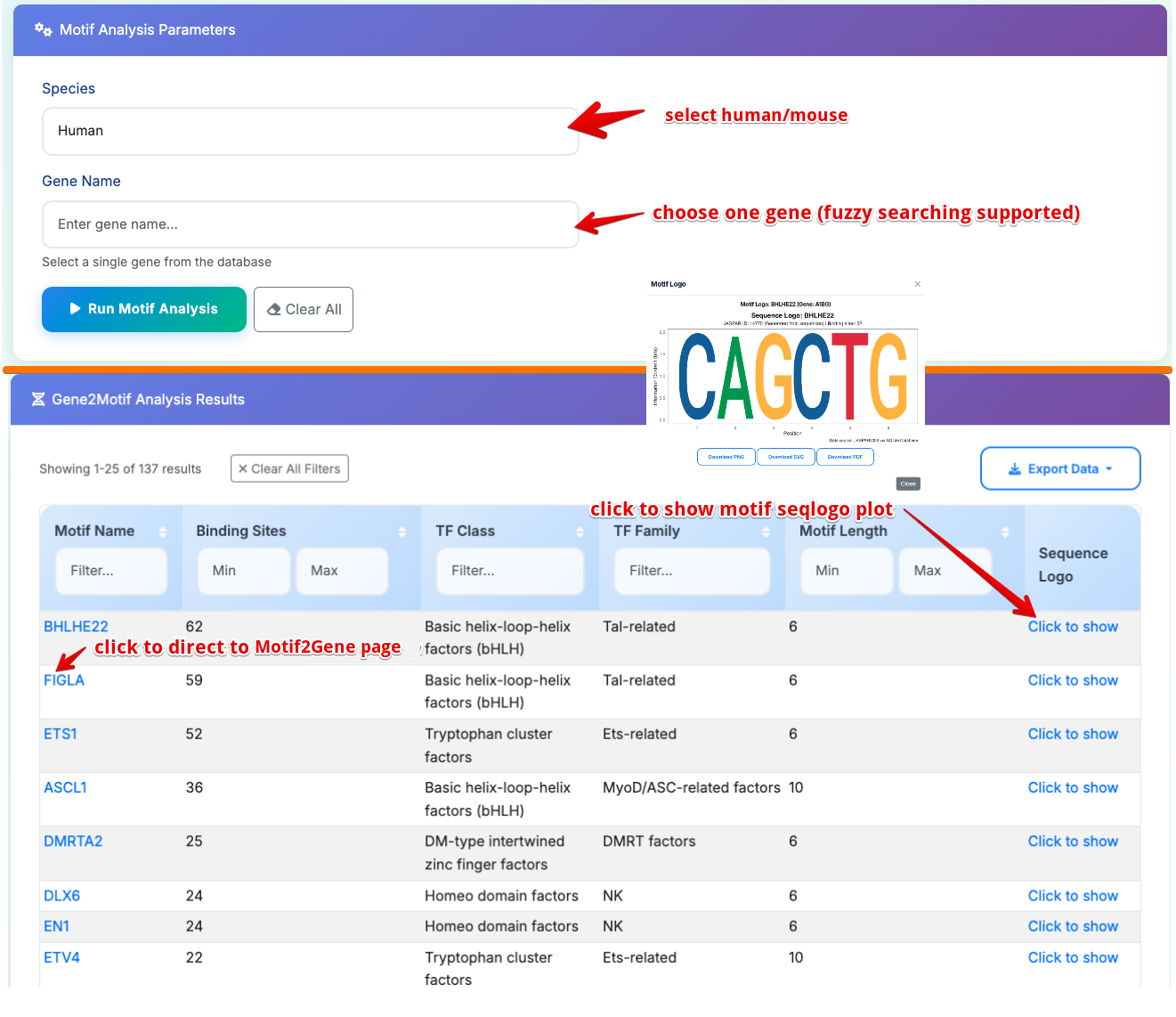
Sub-Module 1: Gene2Motif Analysis
Purpose: Find all transcription factor binding motifs present in a specific gene
Usage:
- Select Species: Choose Human or Mouse from dropdown
- Select Gene: Click dropdown to search and select one gene from database
- Run Analysis: Click Run Motif Analysis to execute
Results Table Columns: - Motif Name: JASPAR motif identifier with cross-reference links - Binding Sites: Number of predicted binding sites in promoter sequence - TF Class: Transcription factor structural classification - TF Family: Transcription factor family grouping - Motif Length: Length of consensus binding sequence - Show Motif Logo: Sequence logo visualization
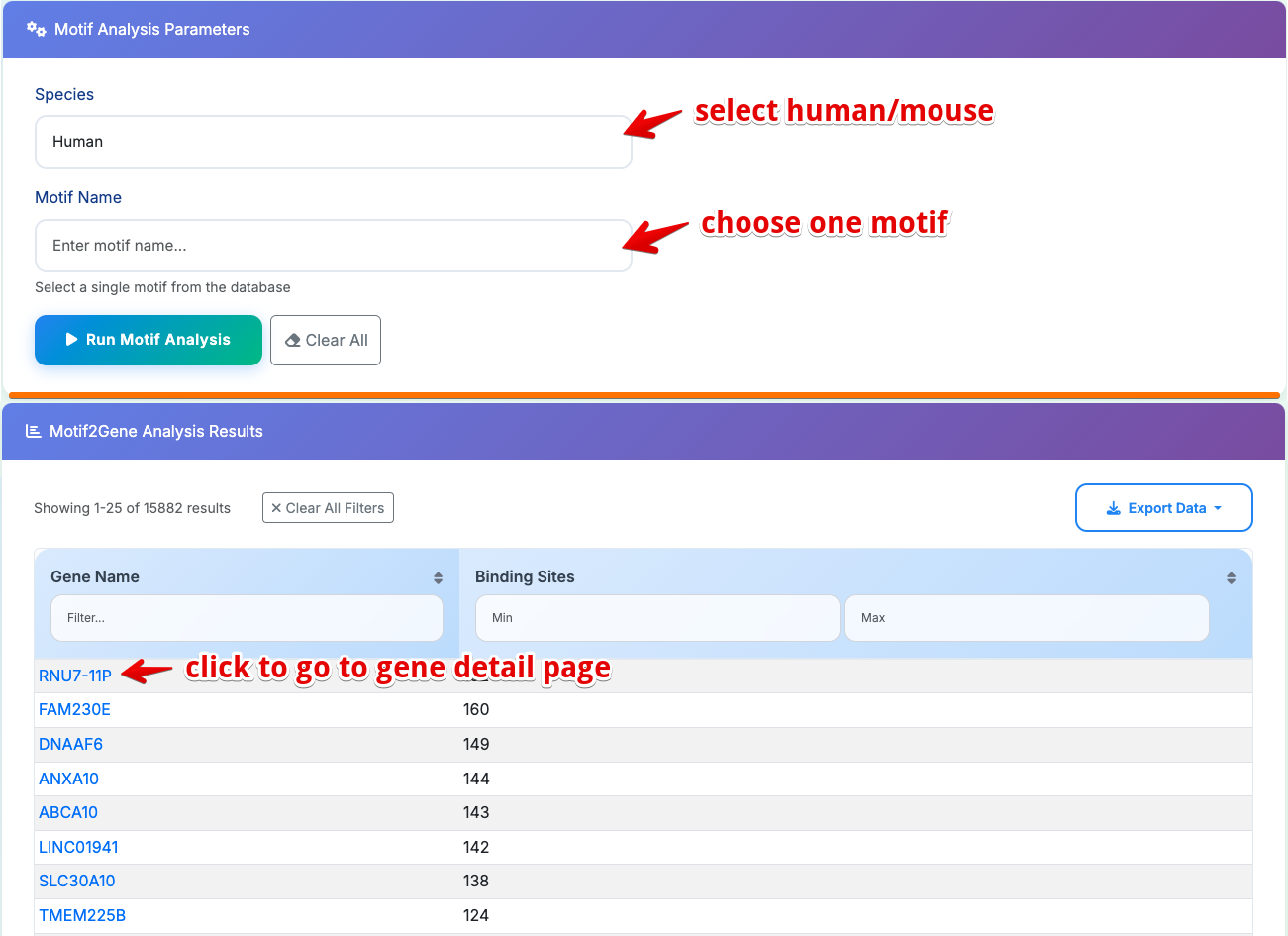
Sub-Module 2: Motif2Gene Analysis
Purpose: Find all genes containing a specific transcription factor binding motif
Usage:
- Select Species: Choose Human or Mouse from dropdown
- Select Motif: Click dropdown to search and select motif from database
- Run Analysis: Click Run Motif Analysis to execute
Results Table Columns:
- Gene Name: Gene symbol with direct links to gene details pages
- Binding Sites: Number of predicted binding sites
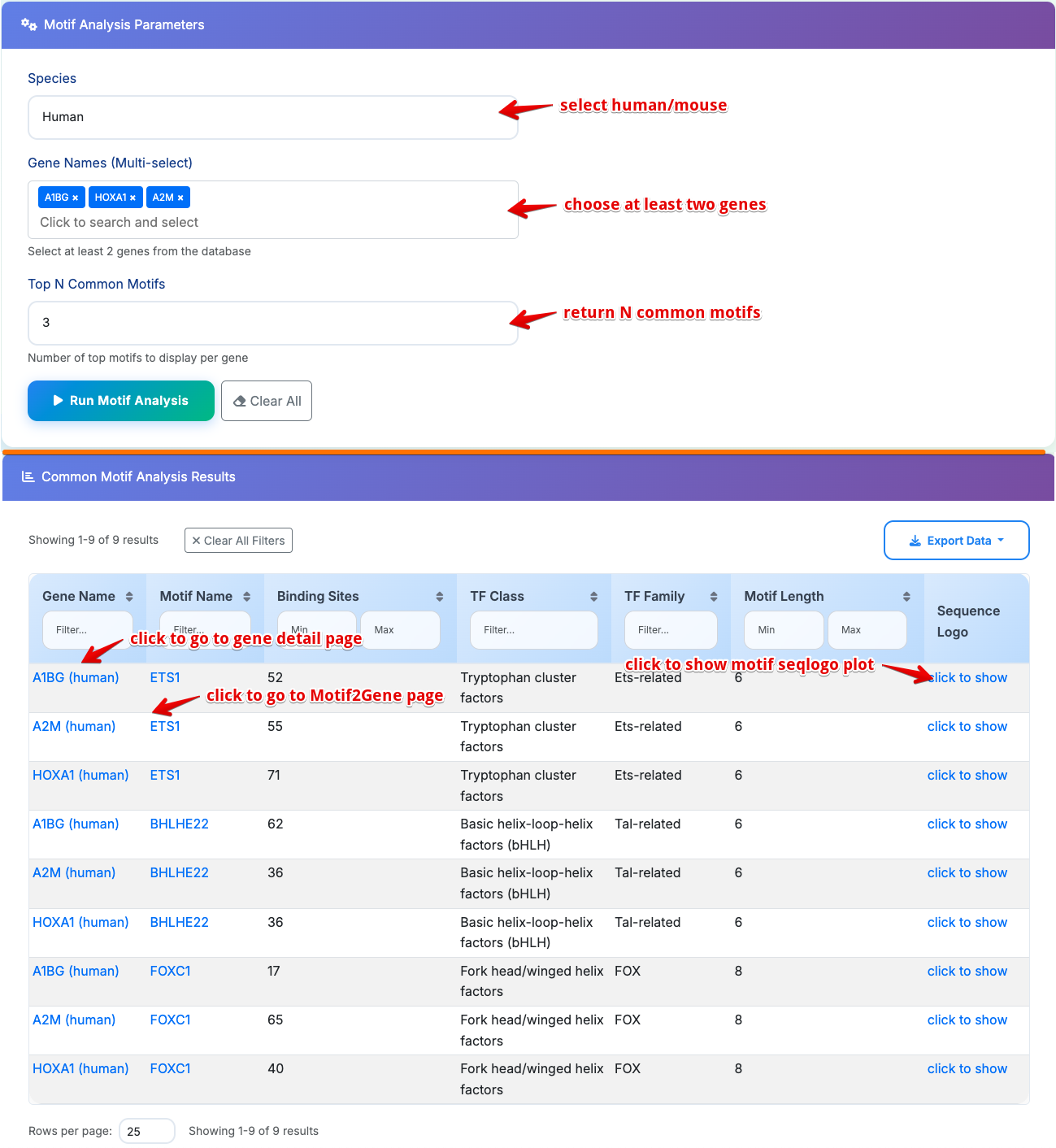
Sub-Module 3: Common Motif Analysis
Purpose: Identify shared transcription factor binding motifs among multiple genes
Usage Steps: 1. Select Species: Choose Human or Mouse from dropdown 2. Multi-Gene Selection: Click dropdown to select multiple genes (minimum 2 required) Selected genes appear as tags with removal options; Supports fuzzy search for gene discovery 3. Set Parameters: Configure "Top N Common Motifs" (5-50, default: 10) 4. Run Analysis: Click Run Motif Analysis to execute
Results Table Columns:
- Gene Name: Gene symbol with species-specific links
- Motif Name: Shared motif identifier with cross-navigation to Motif2Gene
- Binding Sites: Number of binding sites in each gene
- TF Class: Transcription factor classification
- TF Family: Transcription factor family
- Motif Length: Consensus sequence length
- Show Motif Logo: Sequence logo visualization
How to Download Data?
PRIME provides direct access to three categories of pre-compiled datasets without complex filtering or generation processes.
Data Categories:
1. Polycomb Regulatory Targets
- 8 datasets: Human/Mouse × Normal/Disease × All/High-confidence
- File sizes: 1.4MB - 49.4MB
- Content: Cross-species and condition-specific Polycomb regulatory targets
2. Disease Related Targets
- Single Excel file (453KB) with 4 sheets
- Content: Computational analysis results of disease-specific Polycomb targets
3. Literature Mining Results
- Single Excel file (768KB)
- Content: Systematically extracted Polycomb-related information from peer-reviewed literature
API Documentation
PRIME provides a comprehensive RESTful API for accessing all platform data and functionality programmatically. Perfect for researchers building custom analysis pipelines or integrating PRIME data into larger workflows.
Base URL: https://primedb.org/api/
Gene Information APIs
/api/gene/{gene_name}/info- Gene details and annotations/api/gene/{gene_name}/coordinates- Genomic coordinates for browser/api/gene/{gene_name}/external_db- External database links
Expression Data APIs
/api/gene/{gene_name}/bodymap- PTS tissue expression data/api/expression/{atlas}/{gene_name}/plot- Expression atlas plots (GTEx, TCGA, HPA, FANTOM5)
Search & Browse APIs
POST /api/search/execute- Advanced search with filtering/api/gene_suggestions- Gene name autocomplete/api/browse/datasets- Dataset browsing with filters
Analysis APIs
POST /api/analysis/comparison/run- Species/tissue/status comparisonsPOST /api/analysis/network/run- Regulatory network analysisPOST /api/analysis/motif/run- Motif analysis (Gene2Motif, Motif2Gene, Common)
Download APIs
/api/download/polycomb/{dataset}/{format}- Pre-compiled datasetsPOST /api/search/download- Custom search result exports
Python Integration Example
import requests
# Get gene information
response = requests.get("http://primedb.org/api/gene/HOXA1/info?species=human")
gene_data = response.json()
# Search genes
search_data = {
"gene_names": ["HOXA1", "PAX2"],
"species": "human",
"confidence": "high"
}
response = requests.post("http://primedb.org/api/search/execute", json=search_data)
results = response.json()
R Integration Example
library(httr)
library(jsonlite)
# Get gene information
response <- GET("http://primedb.org/api/gene/HOXA1/info?species=human")
gene_data <- fromJSON(content(response, "text"))
# Search genes
search_data <- list(
gene_names = c("HOXA1", "PAX2"),
species = "human",
confidence = "high"
)
response <- POST("http://primedb.org/api/search/execute",
body = search_data,
encode = "json")
results <- fromJSON(content(response, "text"))
Response Format:
All APIs return JSON with standardized error handling:
- HTTP 200: Success
- HTTP 400: Invalid parameters
- HTTP 404: Resource not found
Frequently Asked Questions
How is PRIME built?
Backend Framework:
- Flask web framework with Python for rapid development and API endpoints
- PostgreSQL database for robust data storage and complex queries
- SQLAlchemy ORM for efficient database management and migrations
Data Processing:
- Python-R Hybrid Architecture for optimal performance - Python handles database operations and dynamic interactive visualizations, while R generates static publication-quality plots
- SQLite databases for specialized datasets (motif analysis, expression atlases, literature mining)
- BigWig file format for efficient genomic track storage and IGV.js integration
Frontend Technologies:
- Bootstrap 5 for responsive design and professional UI components
- IGV.js for interactive genome browser functionality
- Plotly.js for dynamic data visualizations and charts
- Custom JavaScript for advanced user interactions and API communication
Scientific Integration:
- Multi-omics data integration from ChIP-seq, RNA-seq, and literature sources
- External database APIs (GTEx, TCGA, HPA, FANTOM5) for comprehensive expression analysis
- R statistical packages for publication-quality static visualizations (expression atlases, network plots, violin plots, motif logos)
- Python libraries for dynamic interactive charts and real-time data visualizations
Deployment:
- Docker containerization for consistent deployment across environments
- Gunicorn WSGI server for production deployment within containers
- ProxyFix middleware for reverse proxy compatibility
- Environment-based configuration for flexible deployment across platforms
This architecture ensures scalable performance, scientific accuracy, and user-friendly interfaces for comprehensive Polycomb research.
Which species are supported?
PRIME currently supports only Human (Homo sapiens) and Mouse (Mus musculus), as Polycomb regulation is best understood, highly conserved, and most thoroughly mapped in these species.
How are confidence levels determined?
- Adaptive tissue-specific thresholds: Different percentile cutoffs adjusted by tissue size to ensure comparable results
- Integrated multi-omics evidence: Combined H3K27me3 ChIP-seq binding and perturbed RNA-seq expression data
- Hierarchical scoring system: Three-tier classification based on final composite scores
Three levels:
- High: Strongest Polycomb regulation evidence (80-90th percentile (depending on tissue size: ≥10,000 genes use 90%, 5,000-10,000 use 85%, <5,000 use 80%))
- Medium: Moderate regulation evidence (>50th percentile threshold)
- Low: Weak or no clear regulatory relationship
Why are some search results empty?
Because there is no corresponding data or no significant results for this gene/tissue combination in our database.
Why might pages load slowly?
PRIME processes large datasets in real-time. Complex analyses may take 30-60 seconds to complete. We recommend patience for best results.
Can I download all data at once?
Yes, use the Download page to get pre-compiled datasets, or export custom results from Search and Browse pages.
How to cite PRIME?
Please cite PRIME in your publications. Citation information will be provided upon database publication.